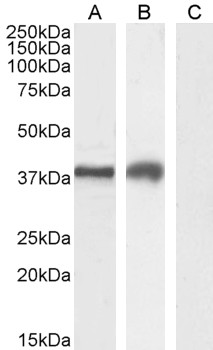
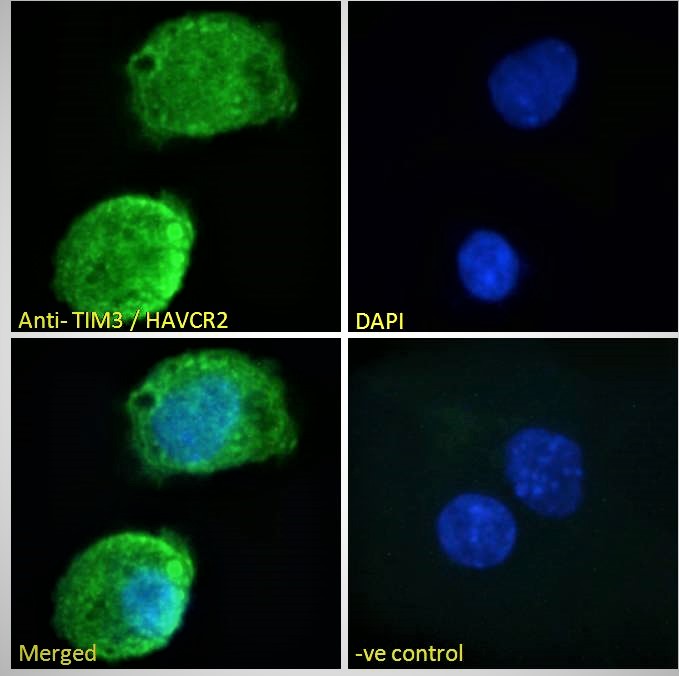
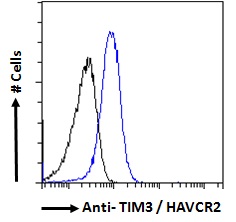
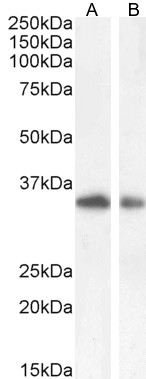
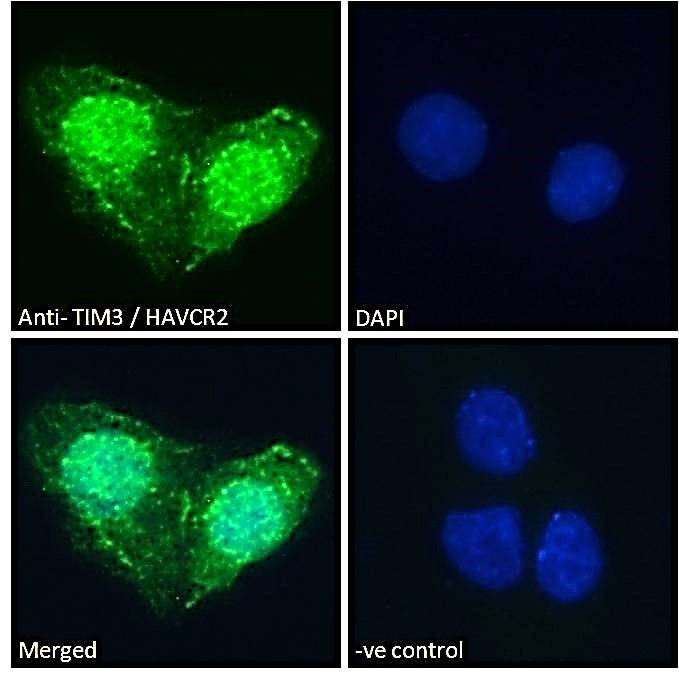
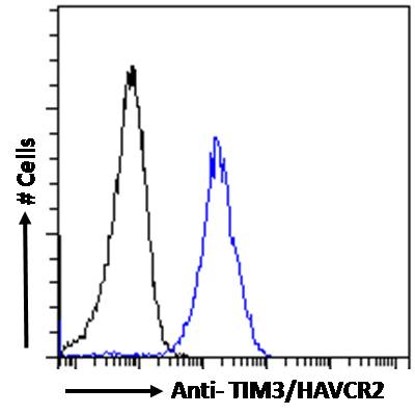
Goat Anti-TIM3 / HAVCR2 Antibody
Peptide-affinity purified goat antibody
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
| WB, IF, FC, Pep-ELISA |
|---|---|
| Primary Accession | Q8TDQ0 |
| Other Accession | NP_116171, 84868 |
| Reactivity | Human |
| Predicted | Dog |
| Host | Goat |
| Clonality | Polyclonal |
| Concentration | 100ug/200ul |
| Isotype | IgG |
| Calculated MW | 33394 Da |
| Gene ID | 84868 |
|---|---|
| Other Names | Hepatitis A virus cellular receptor 2, HAVcr-2, T-cell immunoglobulin and mucin domain-containing protein 3, TIMD-3, T-cell immunoglobulin mucin receptor 3, TIM-3, T-cell membrane protein 3, HAVCR2, TIM3, TIMD3 |
| Format | 0.5 mg IgG/ml in Tris saline (20mM Tris pH7.3, 150mM NaCl), 0.02% sodium azide, with 0.5% bovine serum albumin |
| Storage | Maintain refrigerated at 2-8°C for up to 6 months. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | Goat Anti-TIM3 / HAVCR2 Antibody is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | HAVCR2 |
|---|---|
| Synonyms | TIM3, TIMD3 |
| Function | Cell surface receptor implicated in modulating innate and adaptive immune responses. Generally accepted to have an inhibiting function. Reports on stimulating functions suggest that the activity may be influenced by the cellular context and/or the respective ligand (PubMed:24825777). Regulates macrophage activation (PubMed:11823861). Inhibits T-helper type 1 lymphocyte (Th1)-mediated auto- and alloimmune responses and promotes immunological tolerance (PubMed:14556005). In CD8+ cells attenuates TCR-induced signaling, specifically by blocking NF-kappaB and NFAT promoter activities resulting in the loss of IL-2 secretion. The function may implicate its association with LCK proposed to impair phosphorylation of TCR subunits, and/or LGALS9-dependent recruitment of PTPRC to the immunological synapse (PubMed:24337741, PubMed:26492563). In contrast, shown to activate TCR-induced signaling in T-cells probably implicating ZAP70, LCP2, LCK and FYN (By similarity). Expressed on Treg cells can inhibit Th17 cell responses (PubMed:24838857). Receptor for LGALS9 (PubMed:16286920, PubMed:24337741). Binding to LGALS9 is believed to result in suppression of T-cell responses; the resulting apoptosis of antigen- specific cells may implicate HAVCR2 phosphorylation and disruption of its association with BAG6. Binding to LGALS9 is proposed to be involved in innate immune response to intracellular pathogens. Expressed on Th1 cells interacts with LGALS9 expressed on Mycobacterium tuberculosis- infected macrophages to stimulate antibactericidal activity including IL-1 beta secretion and to restrict intracellular bacterial growth (By similarity). However, the function as receptor for LGALS9 has been challenged (PubMed:23555261). Also reported to enhance CD8+ T-cell responses to an acute infection such as by Listeria monocytogenes (By similarity). Receptor for phosphatidylserine (PtSer); PtSer-binding is calcium-dependent. May recognize PtSer on apoptotic cells leading to their phagocytosis. Mediates the engulfment of apoptotic cells by dendritic cells. Expressed on T-cells, promotes conjugation but not engulfment of apoptotic cells. Expressed on dendritic cells (DCs) positively regulates innate immune response and in synergy with Toll- like receptors promotes secretion of TNF-alpha. In tumor-imfiltrating DCs suppresses nucleic acid-mediated innate immune repsonse by interaction with HMGB1 and interfering with nucleic acid-sensing and trafficking of nucleid acids to endosomes (By similarity). Expressed on natural killer (NK) cells acts as a coreceptor to enhance IFN-gamma production in response to LGALS9 (PubMed:22323453). In contrast, shown to suppress NK cell-mediated cytotoxicity (PubMed:22383801). Negatively regulates NK cell function in LPS-induced endotoxic shock (By similarity). |
| Cellular Location | Membrane; Single-pass type I membrane protein. Cell junction. Cell membrane. Note=Localizes to the immunological synapse between CD8+ T-cells and target cells |
| Tissue Location | Expressed in T-helper type 1 (Th1) lymphocytes. Expressed on regulatory T (Treg) cells after TCR stimulation. Expressed in dendritic cells and natural killer (NK) cells. Expressed in epithelial tissues. Expression is increased on CD4+ and CD8+ T-cells in chronic hepatitis C virus (HCV) infection. In progressive HIV-1 infection, expression is up-regulated on HIV-1-specific CD8 T-cells |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
Background
CD4 (MIM 186940)-positive T helper lymphocytes can be divided into types 1 (Th1) and 2 (Th2) on the basis of their cytokine secretion patterns. Th1 cells and their associated cytokines are involved in cell-mediated immunity to intracellular pathogens and delayed-type hypersensitivity reactions, whereas Th2 cells are involved in the control of extracellular helminthic infections and the promotion of atopic and allergic diseases. The 2 types of cells also cross-regulate the functions of the other. TIM3 is a Th1-specific cell surface protein that regulates macrophage activation and enhances the severity of experimental autoimmune encephalomyelitis in mice.
References
Genetic variations and haplotypes in TIM-3 gene and the risk of gastric cancer. Cao B, et al. Cancer Immunol Immunother, 2010 Sep 2. PMID 20811886.
Variation at the NFATC2 Locus Increases the Risk of Thiazolinedinedione-Induced Edema in the Diabetes REduction Assessment with ramipril and rosiglitazone Medication (DREAM) Study. Bailey SD, et al. Diabetes Care, 2010 Jul 13. PMID 20628086.
Tim-3/galectin-9 pathway: regulation of Th1 immunity through promotion of CD11b+Ly-6G+ myeloid cells. Dardalhon V, et al. J Immunol, 2010 Aug 1. PMID 20574007.
Interleukin-9 polymorphism in infants with respiratory syncytial virus infection: an opposite effect in boys and girls. Schuurhof A, et al. Pediatr Pulmonol, 2010 Jun. PMID 20503287.
Large-scale candidate gene analysis of spontaneous clearance of hepatitis C virus. Mosbruger TL, et al. J Infect Dis, 2010 May 1. PMID 20331378.
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.













 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.