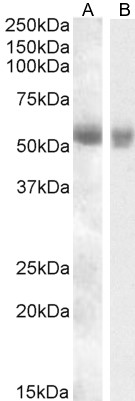
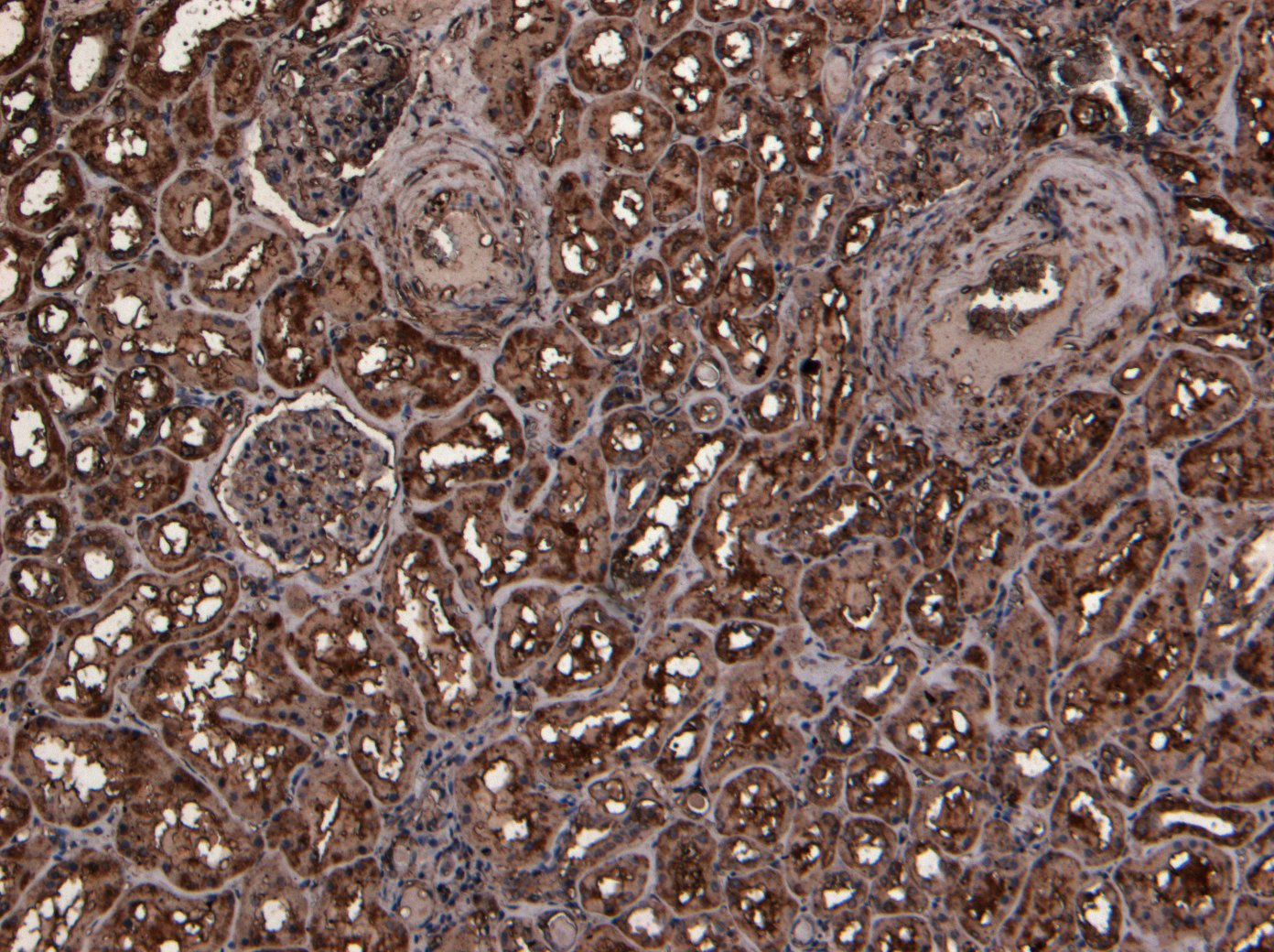
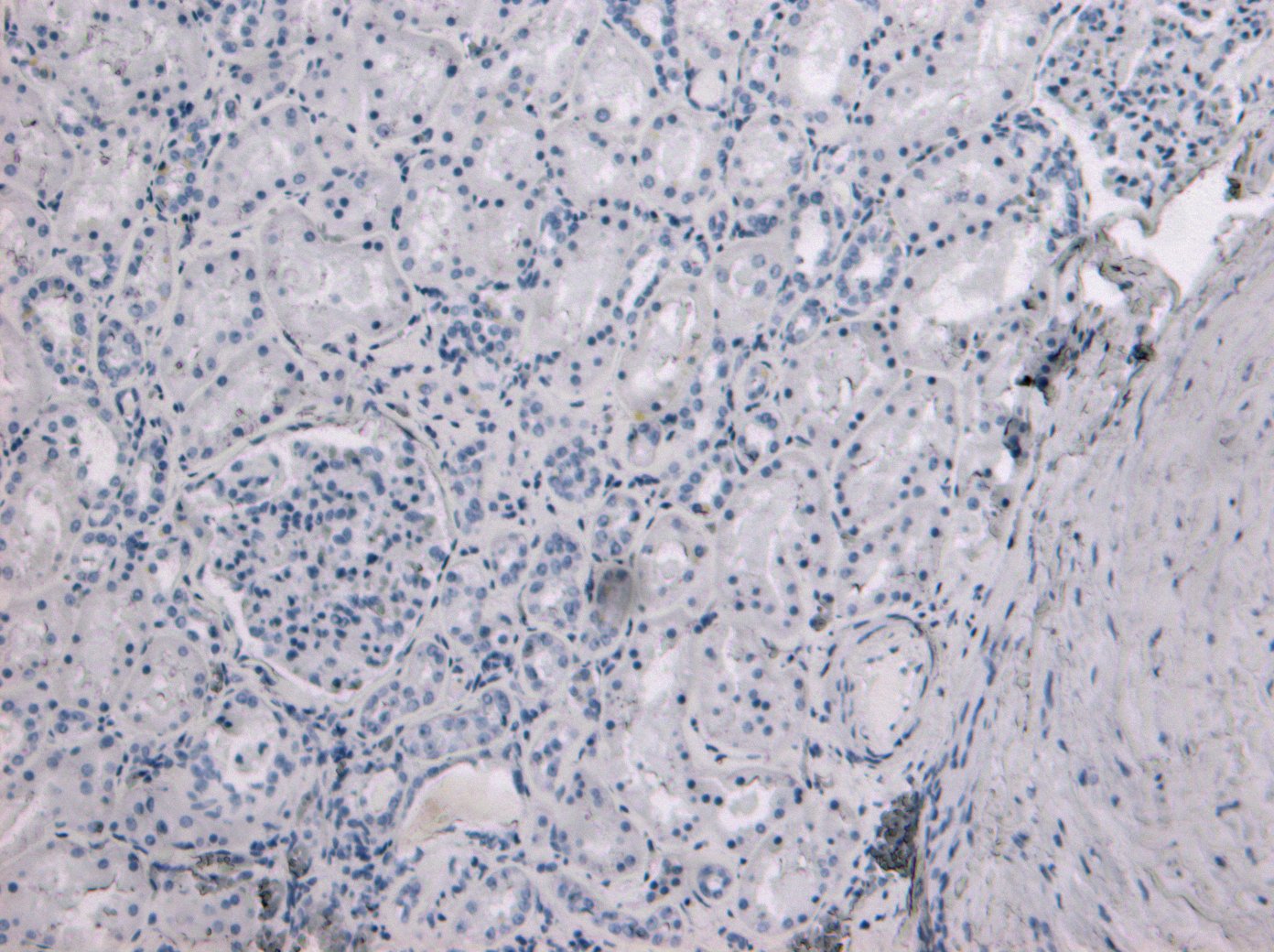
Goat Anti-TMPRSS2 Antibody
Peptide-affinity purified goat antibody
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
| WB, IHC, Pep-ELISA |
|---|---|
| Primary Accession | O15393 |
| Other Accession | NP_005647, 7113 |
| Reactivity | Human |
| Host | Goat |
| Clonality | Polyclonal |
| Concentration | 100ug/200ul |
| Isotype | IgG |
| Calculated MW | 53859 Da |
| Gene ID | 7113 |
|---|---|
| Other Names | Transmembrane protease serine 2, 3.4.21.-, Serine protease 10, Transmembrane protease serine 2 non-catalytic chain, Transmembrane protease serine 2 catalytic chain, TMPRSS2, PRSS10 |
| Format | 0.5 mg IgG/ml in Tris saline (20mM Tris pH7.3, 150mM NaCl), 0.02% sodium azide, with 0.5% bovine serum albumin |
| Storage | Maintain refrigerated at 2-8°C for up to 6 months. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | Goat Anti-TMPRSS2 Antibody is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | TMPRSS2 (HGNC:11876) |
|---|---|
| Synonyms | PRSS10 |
| Function | Plasma membrane-anchored serine protease that cleaves at arginine residues (PubMed:32703818). Participates in proteolytic cascades of relevance for the normal physiologic function of the prostate (PubMed:25122198). Androgen-induced TMPRSS2 activates several substrates that include pro-hepatocyte growth factor/HGF, the protease activated receptor-2/F2RL1 or matriptase/ST14 leading to extracellular matrix disruption and metastasis of prostate cancer cells (PubMed:15537383, PubMed:26018085, PubMed:25122198). In addition, activates trigeminal neurons and contribute to both spontaneous pain and mechanical allodynia (By similarity). |
| Cellular Location | Cell membrane; Single-pass type II membrane protein |
| Tissue Location | Expressed in several tissues that comprise large populations of epithelial cells with the highest level of transcripts measured in the prostate gland. Expressed in type II pneumocytes in the lung (at protein level). Expressed strongly in small intestine. Also expressed in colon, stomach and salivary gland. Coexpressed with ACE2 within lung type II pneumocytes, ileal absorptive enterocytes, intestinal epithelial cells, cornea, gallbladder and nasal goblet secretory cells (Ref.21). {ECO:0000269|PubMed:11169526, ECO:0000269|PubMed:20382709, ECO:0000269|PubMed:21325420, ECO:0000269|PubMed:32404436, ECO:0000269|Ref.21} |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
Background
This gene encodes a protein that belongs to the serine protease family. The encoded protein contains a type II transmembrane domain, a receptor class A domain, a scavenger receptor cysteine-rich domain and a protease domain. Serine proteases are known to be involved in many physiological and pathological processes. This gene was demonstrated to be up-regulated by androgenic hormones in prostate cancer cells and down-regulated in androgen-independent prostate cancer tissue. The protease domain of this protein is thought to be cleaved and secreted into cell media after autocleavage. Alternatively spliced transcript variants encoding different isoforms have been found for this gene.
References
Association of SPINK1 expression and TMPRSS2:ERG fusion with prognosis in endocrine-treated prostate cancer. Leinonen KA, et al. Clin Cancer Res, 2010 May 15. PMID 20442300.
Prevalence of TMPRSS2-ERG and SLC45A3-ERG gene fusions in a large prostatectomy cohort. Esgueva R, et al. Mod Pathol, 2010 Apr. PMID 20118910.
Prostate cancer genes associated with TMPRSS2-ERG gene fusion and prognostic of biochemical recurrence in multiple cohorts. Barwick BG, et al. Br J Cancer, 2010 Feb 2. PMID 20068566.
Induced chromosomal proximity and gene fusions in prostate cancer. Mani RS, et al. Science, 2009 Nov 27. PMID 19933109.
Functional screening of FxxLF-like peptide motifs identifies SMARCD1/BAF60a as an androgen receptor cofactor that modulates TMPRSS2 expression. van de Wijngaart DJ, et al. Mol Endocrinol, 2009 Nov. PMID 19762545.
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.













 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.