beta-Catenin (p120) Antibody - With BSA and Azide
Rabbit Polyclonal Antibody
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
| WB, IHC-P, IF, FC |
|---|---|
| Primary Accession | P35222 |
| Other Accession | 1499, 476018 |
| Reactivity | Human, Mouse |
| Host | Rabbit |
| Clonality | Polyclonal |
| Isotype | Rabbit / IgG |
| Clone Names | |
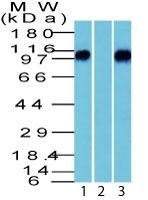
| Calculated MW | 92kDa |
| Gene ID | 1499 |
|---|---|
| Other Names | Catenin beta-1, Beta-catenin, CTNNB1, CTNNB |
| Format | 200ug/ml of Ab purified from rabbit anti-serum by Protein A. Prepared in 10mM PBS with 0.05% BSA & 0.05% azide. Also available WITHOUT BSA at 1.0mg/ml. |
| Storage | Store at 2 to 8°C.Antibody is stable for 24 months. |
| Precautions | beta-Catenin (p120) Antibody - With BSA and Azide is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | CTNNB1 (HGNC:2514) |
|---|---|
| Synonyms | CTNNB |
| Function | Key downstream component of the canonical Wnt signaling pathway (PubMed:17524503, PubMed:18077326, PubMed:18086858, PubMed:18957423, PubMed:21262353, PubMed:22155184, PubMed:22647378, PubMed:22699938). In the absence of Wnt, forms a complex with AXIN1, AXIN2, APC, CSNK1A1 and GSK3B that promotes phosphorylation on N- terminal Ser and Thr residues and ubiquitination of CTNNB1 via BTRC and its subsequent degradation by the proteasome (PubMed:17524503, PubMed:18077326, PubMed:18086858, PubMed:18957423, PubMed:21262353, PubMed:22155184, PubMed:22647378, PubMed:22699938). In the presence of Wnt ligand, CTNNB1 is not ubiquitinated and accumulates in the nucleus, where it acts as a coactivator for transcription factors of the TCF/LEF family, leading to activate Wnt responsive genes (PubMed:17524503, PubMed:18077326, PubMed:18086858, PubMed:18957423, PubMed:21262353, PubMed:22155184, PubMed:22647378, PubMed:22699938). Involved in the regulation of cell adhesion, as component of an E-cadherin:catenin adhesion complex (By similarity). Acts as a negative regulator of centrosome cohesion (PubMed:18086858). Involved in the CDK2/PTPN6/CTNNB1/CEACAM1 pathway of insulin internalization (PubMed:21262353). Blocks anoikis of malignant kidney and intestinal epithelial cells and promotes their anchorage-independent growth by down-regulating DAPK2 (PubMed:18957423). Disrupts PML function and PML- NB formation by inhibiting RANBP2-mediated sumoylation of PML (PubMed:22155184). Promotes neurogenesis by maintaining sympathetic neuroblasts within the cell cycle (By similarity). Involved in chondrocyte differentiation via interaction with SOX9: SOX9-binding competes with the binding sites of TCF/LEF within CTNNB1, thereby inhibiting the Wnt signaling (By similarity). Acts as a positive regulator of odontoblast differentiation during mesenchymal tooth germ formation, via promoting the transcription of differentiation factors such as LEF1, BMP2 and BMP4 (By similarity). Activity is repressed in a MSX1-mediated manner at the bell stage of mesenchymal tooth germ formation which prevents premature differentiation of odontoblasts (By similarity). |
| Cellular Location | Cytoplasm. Nucleus. Cytoplasm, cytoskeleton {ECO:0000250|UniProtKB:B6V8E6}. Cell junction, adherens junction. Cell junction {ECO:0000250|UniProtKB:B6V8E6}. Cell membrane. Cytoplasm, cytoskeleton, microtubule organizing center, centrosome. Cytoplasm, cytoskeleton, spindle pole. Synapse {ECO:0000250|UniProtKB:Q02248} Cytoplasm, cytoskeleton, cilium basal body {ECO:0000250|UniProtKB:Q02248}. Note=Colocalized with RAPGEF2 and TJP1 at cell-cell contacts (By similarity). Cytoplasmic when it is un-stable (highly phosphorylated) or bound to CDH1. Translocates to the nucleus when it is stabilized (low level of phosphorylation). Interaction with GLIS2 and MUC1 promotes nuclear translocation. Interaction with EMD inhibits nuclear localization. The majority of beta-catenin is localized to the cell membrane. In interphase, colocalizes with CROCC between CEP250 puncta at the proximal end of centrioles, and this localization is dependent on CROCC and CEP250. In mitosis, when NEK2 activity increases, it localizes to centrosomes at spindle poles independent of CROCC. Colocalizes with CDK5 in the cell-cell contacts and plasma membrane of undifferentiated and differentiated neuroblastoma cells. Interaction with FAM53B promotes translocation to the nucleus (PubMed:25183871). {ECO:0000250|UniProtKB:B6V8E6, ECO:0000269|PubMed:25183871} |
| Tissue Location | Expressed in several hair follicle cell types: basal and peripheral matrix cells, and cells of the outer and inner root sheaths. Expressed in colon. Present in cortical neurons (at protein level). Expressed in breast cancer tissues (at protein level) (PubMed:29367600). |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
Background
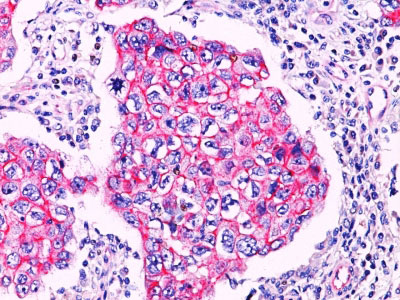
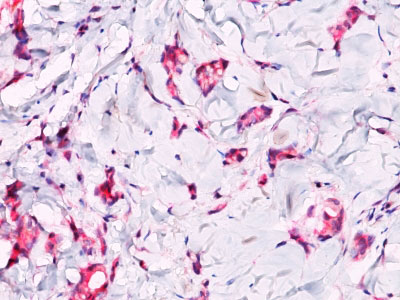
Beta-catenin associates with the cytoplasmic portion of E-cadherin, which is necessary for the function of E-cadherin as an adhesion molecule. In normal tissues, beta-catenin is localized to the membrane of epithelial cells, consistent with its role in the cell adhesion complex. In breast ductal neoplasia, beta-catenin is usually localized in cellular membranes. However, in lobular neoplasia, a marked redistribution of beta-catenin throughout the cytoplasm results in a diffuse cytoplasmic pattern. Immuno-staining of beta-catenin and E-cadherin is helps in the accurate identification of ductal and lobular neoplasms, including a distinction between low-grade ductal carcinoma in situ (DCIS) and lobular carcinoma. Additionally, some rectal and gastric adenocarcinomas demonstrate diffuse cytoplasmic beta-catenin staining and a lack of membranous staining, mimicking the staining pattern observed with lobular breast carcinomas.
References
Dabbs DJ et. al. Am J Surg Path. 2007;31:427-437. | Sarrio D et. al. Oncogene. 2004;23:3272-3283. | Mastracci TL et. al. Mod Path. 2005;18:741-751
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.













 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.