KEAP1 Antibody
Purified Mouse Monoclonal Antibody
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
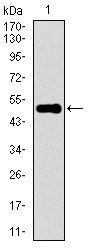
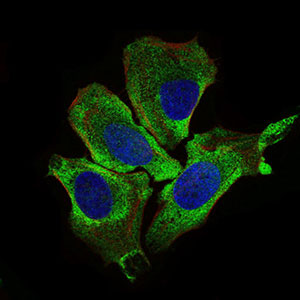
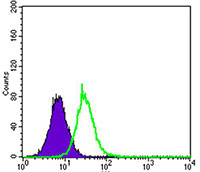
| WB, FC, ICC, E |
|---|---|
| Primary Accession | Q14145 |
| Reactivity | Human, Mouse |
| Host | Mouse |
| Clonality | Monoclonal |
| Clone Names | 1F10B6 |
| Isotype | IgG1 |
| Calculated MW | 69.7kDa |
| Description | This gene encodes a protein containing KELCH-1 like domains, as well as a BTB/POZ domain. Kelch-like ECH-associated protein 1 interacts with NF-E2-related factor 2 in a redox-sensitive manner and the dissociation of the proteins in the cytoplasm is followed by transportation of NF-E2-related factor 2 to the nucleus. This interaction results in the expression of the catalytic subunit of gamma-glutamylcysteine synthetase. Two alternatively spliced transcript variants encoding the same isoform have been found for this gene. |
| Immunogen | Purified recombinant fragment of human KEAP1 (AA: 380-624) expressed in E. Coli. |
| Formulation | Ascitic fluid containing 0.03% sodium azide. |
| Gene ID | 9817 |
|---|---|
| Other Names | Kelch-like ECH-associated protein 1, Cytosolic inhibitor of Nrf2, INrf2, Kelch-like protein 19, KEAP1, INRF2, KIAA0132, KLHL19 |
| Dilution | E~~1/10000 WB~~1/500 - 1/2000 IF~~1/50 FC~~1/200 - 1/400 |
| Storage | Maintain refrigerated at 2-8°C for up to 6 months. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | KEAP1 Antibody is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | KEAP1 {ECO:0000303|PubMed:14585973, ECO:0000312|HGNC:HGNC:23177} |
|---|---|
| Function | Substrate-specific adapter of a BCR (BTB-CUL3-RBX1) E3 ubiquitin ligase complex that regulates the response to oxidative stress by targeting NFE2L2/NRF2 for ubiquitination (PubMed:14585973, PubMed:15379550, PubMed:15572695, PubMed:15983046, PubMed:15601839, PubMed:37339955). KEAP1 acts as a key sensor of oxidative and electrophilic stress: in normal conditions, the BCR(KEAP1) complex mediates ubiquitination and degradation of NFE2L2/NRF2, a transcription factor regulating expression of many cytoprotective genes (PubMed:15601839, PubMed:16006525). In response to oxidative stress, different electrophile metabolites trigger non-enzymatic covalent modifications of highly reactive cysteine residues in KEAP1, leading to inactivate the ubiquitin ligase activity of the BCR(KEAP1) complex, promoting NFE2L2/NRF2 nuclear accumulation and expression of phase II detoxifying enzymes (PubMed:19489739, PubMed:16006525, PubMed:17127771, PubMed:18251510, PubMed:29590092). In response to selective autophagy, KEAP1 is sequestered in inclusion bodies following its interaction with SQSTM1/p62, leading to inactivation of the BCR(KEAP1) complex and activation of NFE2L2/NRF2 (PubMed:20452972). The BCR(KEAP1) complex also mediates ubiquitination of SQSTM1/p62, increasing SQSTM1/p62 sequestering activity and degradation (PubMed:28380357). The BCR(KEAP1) complex also targets BPTF and PGAM5 for ubiquitination and degradation by the proteasome (PubMed:15379550, PubMed:17046835). |
| Cellular Location | Cytoplasm. Nucleus. Note=Mainly cytoplasmic (PubMed:15601839). In response to selective autophagy, relocalizes to inclusion bodies following interaction with SQSTM1/p62 (PubMed:20452972). |
| Tissue Location | Broadly expressed, with highest levels in skeletal muscle. |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
References
1.Cell Signal. 2010 Nov;22(11):1645-54. 2.Mol Cancer Ther. 2010 Feb;9(2):336-46.
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.













 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.