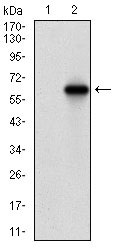
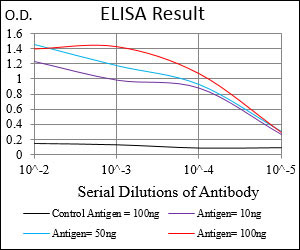
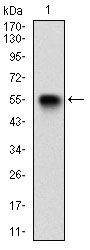
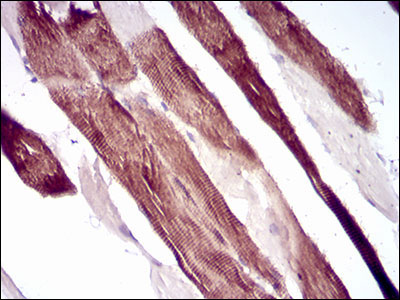
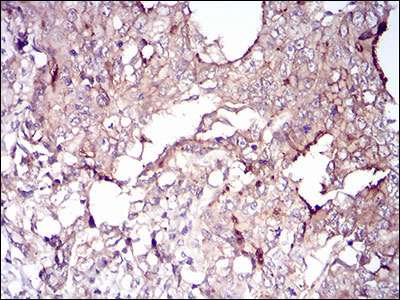
VIL1 Antibody
Purified Mouse Monoclonal Antibody
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
| WB, IHC, E |
|---|---|
| Primary Accession | P09327 |
| Reactivity | Human |
| Host | Mouse |
| Clonality | Monoclonal |
| Clone Names | 5E3B2 |
| Isotype | IgG2b |
| Calculated MW | 92.7kDa |
| Description | This gene encodes a member of a family of calcium-regulated actin-binding proteins. This protein represents a dominant part of the brush border cytoskeleton which functions in the capping, severing, and bundling of actin filaments. Two mRNAs of 2.7 kb and 3.5 kb have been observed; they result from utilization of alternate poly-adenylation signals present in the terminal exon. |
| Immunogen | Purified recombinant fragment of human VIL1 (AA: 1-209) expressed in E. Coli. |
| Formulation | Purified antibody in PBS with 0.05% sodium azide. |
| Gene ID | 7429 |
|---|---|
| Other Names | Villin-1, VIL1, VIL |
| Dilution | E~~1/10000 WB~~1/500 - 1/2000 IHC~~1/200 - 1/1000 |
| Storage | Maintain refrigerated at 2-8°C for up to 6 months. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | VIL1 Antibody is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | VIL1 |
|---|---|
| Synonyms | VIL |
| Function | Epithelial cell-specific Ca(2+)-regulated actin-modifying protein that modulates the reorganization of microvillar actin filaments. Plays a role in the actin nucleation, actin filament bundle assembly, actin filament capping and severing. Binds phosphatidylinositol 4,5-bisphosphate (PIP2) and lysophosphatidic acid (LPA); binds LPA with higher affinity than PIP2. Binding to LPA increases its phosphorylation by SRC and inhibits all actin-modifying activities. Binding to PIP2 inhibits actin-capping and -severing activities but enhances actin-bundling activity. Regulates the intestinal epithelial cell morphology, cell invasion, cell migration and apoptosis. Protects against apoptosis induced by dextran sodium sulfate (DSS) in the gastrointestinal epithelium. Appears to regulate cell death by maintaining mitochondrial integrity. Enhances hepatocyte growth factor (HGF)-induced epithelial cell motility, chemotaxis and wound repair. Upon S.flexneri cell infection, its actin-severing activity enhances actin-based motility of the bacteria and plays a role during the dissemination. |
| Cellular Location | Cytoplasm, cytoskeleton. Cell projection, lamellipodium. Cell projection, ruffle. Cell projection, microvillus Cell projection, filopodium tip. Cell projection, filopodium. Note=Relocalized in the tip of cellular protrusions and filipodial extensions upon infection with S.flexneri in primary intestinal epithelial cells (IEC) and in the tail-like structures forming the actin comets of S.flexneri. Redistributed to the leading edge of hepatocyte growth factor (HGF)-induced lamellipodia (By similarity). Rapidly redistributed to ruffles and lamellipodia structures in response to autotaxin, lysophosphatidic acid (LPA) and epidermal growth factor (EGF) treatment. |
| Tissue Location | Specifically expressed in epithelial cells. Major component of microvilli of intestinal epithelial cells and kidney proximal tubule cells. Expressed in canalicular microvilli of hepatocytes (at protein level). |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
Background
This protein belongs to the aldehyde dehydrogenase family of proteins. Aldehyde dehydrogenase is the second enzyme of the major oxidative pathway of alcohol metabolism. Two major liver isoforms of aldehyde dehydrogenase, cytosolic and mitochondrial, can be distinguished by their electrophoretic mobilities, kinetic properties, and subcellular localizations. Most Caucasians have two major isozymes, while approximately 50% of Orientals have the cytosolic isozyme but not the mitochondrial isozyme. A remarkably higher frequency of acute alcohol intoxication among Orientals than among Caucasians could be related to the absence of a catalytically active form of the mitochondrial isozyme. The increased exposure to acetaldehyde in individuals with the catalytically inactive form may also confer greater susceptibility to many types of cancer. This gene encodes a mitochondrial isoform, which has a low Km for acetaldehydes, and is localized in mitochondrial matrix. Alternative splicing results in multiple transcript variants encoding distinct isoforms. ; ; ;
References
1. Cancer Biol Ther. 2011 Aug 1;12(3):181-90.2. Cancer Biol Ther. 2009 Jun;8(12):1146-53.
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.













 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.