XRCC6 Antibody
Purified Mouse Monoclonal Antibody
- SPECIFICATION
- CITATIONS: 1
- PROTOCOLS
- BACKGROUND

Application
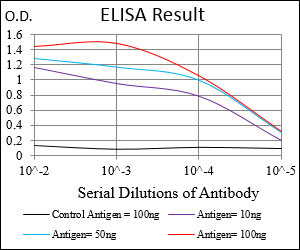
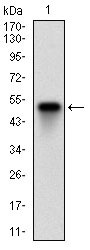
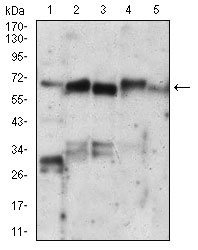
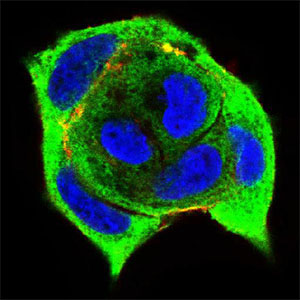
| WB, FC, ICC, E |
|---|---|
| Primary Accession | P12956 |
| Reactivity | Human |
| Host | Mouse |
| Clonality | Monoclonal |
| Clone Names | 7A9E7 |
| Isotype | IgG1 |
| Calculated MW | 69.8kDa |
| Description | The p70/p80 autoantigen is a nuclear complex consisting of two subunits with molecular masses of approximately 70 and 80 kDa. The complex functions as a single-stranded DNA-dependent ATP-dependent helicase. The complex may be involved in the repair of nonhomologous DNA ends such as that required for double-strand break repair, transposition, and V(D)J recombination. High levels of autoantibodies to p70 and p80 have been found in some patients with systemic lupus erythematosus. |
| Immunogen | Purified recombinant fragment of human XRCC6 (AA: 6-214) expressed in E. Coli. |
| Formulation | Purified antibody in PBS with 0.05% sodium azide. |
| Gene ID | 2547 |
|---|---|
| Other Names | X-ray repair cross-complementing protein 6, 3.6.4.-, 4.2.99.-, 5'-deoxyribose-5-phosphate lyase Ku70, 5'-dRP lyase Ku70, 70 kDa subunit of Ku antigen, ATP-dependent DNA helicase 2 subunit 1, ATP-dependent DNA helicase II 70 kDa subunit, CTC box-binding factor 75 kDa subunit, CTC75, CTCBF, DNA repair protein XRCC6, Lupus Ku autoantigen protein p70, Ku70, Thyroid-lupus autoantigen, TLAA, X-ray repair complementing defective repair in Chinese hamster cells 6, XRCC6, G22P1 |
| Dilution | E~~1/10000 WB~~1/500 - 1/2000 IF~~1/200 - 1/1000 FC~~1/200 - 1/400 |
| Storage | Maintain refrigerated at 2-8°C for up to 6 months. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | XRCC6 Antibody is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | XRCC6 |
|---|---|
| Synonyms | G22P1 |
| Function | Single-stranded DNA-dependent ATP-dependent helicase that plays a key role in DNA non-homologous end joining (NHEJ) by recruiting DNA-PK to DNA (PubMed:7957065, PubMed:8621488, PubMed:12145306, PubMed:11493912, PubMed:20493174, PubMed:2466842, PubMed:9742108). Required for double-strand break repair and V(D)J recombination (PubMed:7957065, PubMed:8621488, PubMed:12145306, PubMed:11493912, PubMed:20493174, PubMed:2466842, PubMed:9742108). Also has a role in chromosome translocation (PubMed:7957065, PubMed:8621488, PubMed:12145306, PubMed:11493912, PubMed:20493174, PubMed:2466842, PubMed:9742108). Has a role in chromosome translocation (PubMed:7957065, PubMed:20493174, PubMed:2466842, PubMed:9742108, PubMed:8621488, PubMed:12145306, PubMed:11493912). The DNA helicase II complex binds preferentially to fork-like ends of double-stranded DNA in a cell cycle-dependent manner (PubMed:7957065, PubMed:8621488, PubMed:20493174, PubMed:2466842, PubMed:9742108, PubMed:12145306, PubMed:11493912). It works in the 3'-5' direction (PubMed:20493174, PubMed:2466842, PubMed:9742108, PubMed:7957065, PubMed:8621488, PubMed:12145306, PubMed:11493912). During NHEJ, the XRCC5-XRRC6 dimer performs the recognition step: it recognizes and binds to the broken ends of the DNA and protects them from further resection (PubMed:7957065, PubMed:8621488, PubMed:20493174, PubMed:2466842, PubMed:9742108, PubMed:12145306, PubMed:11493912). Binding to DNA may be mediated by XRCC6 (PubMed:20493174, PubMed:2466842, PubMed:9742108, PubMed:7957065, PubMed:8621488, PubMed:12145306, PubMed:11493912). The XRCC5-XRRC6 dimer acts as a regulatory subunit of the DNA-dependent protein kinase complex DNA-PK by increasing the affinity of the catalytic subunit PRKDC to DNA by 100-fold (PubMed:7957065, PubMed:8621488, PubMed:12145306, PubMed:11493912, PubMed:20493174, PubMed:2466842, PubMed:9742108). The XRCC5-XRRC6 dimer is probably involved in stabilizing broken DNA ends and bringing them together (PubMed:7957065, PubMed:8621488, PubMed:12145306, PubMed:11493912, PubMed:20493174, PubMed:2466842, PubMed:9742108). The assembly of the DNA-PK complex to DNA ends is required for the NHEJ ligation step (PubMed:7957065, PubMed:8621488, PubMed:12145306, PubMed:11493912, PubMed:20493174, PubMed:2466842, PubMed:9742108). Probably also acts as a 5'-deoxyribose-5-phosphate lyase (5'-dRP lyase), by catalyzing the beta-elimination of the 5' deoxyribose-5-phosphate at an abasic site near double-strand breaks (PubMed:20383123). 5'-dRP lyase activity allows to 'clean' the termini of abasic sites, a class of nucleotide damage commonly associated with strand breaks, before such broken ends can be joined (PubMed:20383123). The XRCC5-XRRC6 dimer together with APEX1 acts as a negative regulator of transcription (PubMed:8621488). In association with NAA15, the XRCC5-XRRC6 dimer binds to the osteocalcin promoter and activates osteocalcin expression (PubMed:12145306). Plays a role in the regulation of DNA virus-mediated innate immune response by assembling into the HDP-RNP complex, a complex that serves as a platform for IRF3 phosphorylation and subsequent innate immune response activation through the cGAS-STING pathway (PubMed:28712728). |
| Cellular Location | Nucleus. Chromosome |

Provided below are standard protocols that you may find useful for product applications.
Background
This gene is expressed ubiquitously with higher levels in fetal than in adult tissues. It encodes a protein sharing 93% sequence identity with the mouse protein. Wolf-Hirschhorn syndrome (WHS) is a malformation syndrome associated with a hemizygous deletion of the distal short arm of chromosome 4. This gene is mapped to the 165 kb WHS critical region, and may play a role in the phenotype of the WHS or Pitt-Rogers-Danks syndrome. The encoded protein is found to be capable of reacting with HLA-A2-restricted and tumor-specific cytotoxic T lymphocytes, suggesting a target for use in specific immunotherapy for a large number of cancer patients. This protein has also been shown to be a member of the NELF (negative elongation factor) protein complex that participates in the regulation of RNA polymerase II transcription elongation. ; ;
References
1. Clin Cancer Res. 2013 Mar 15;19(6):1547-56.2. Mol Carcinog. 2012 Oct;51 Suppl 1:E183-90.
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.














 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.