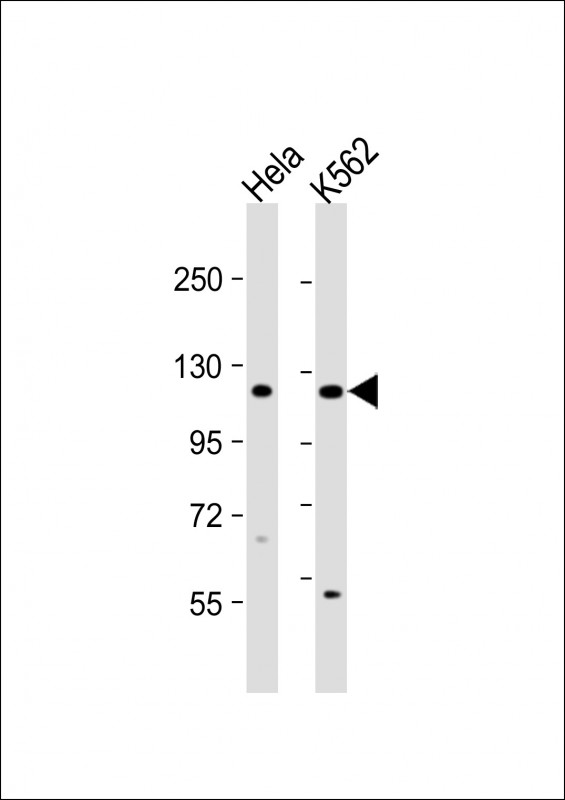
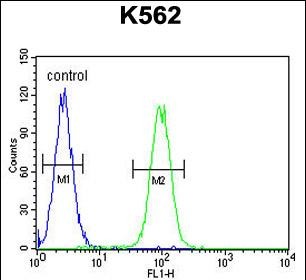
DDX11 Antibody (C-term)
Affinity Purified Rabbit Polyclonal Antibody (Pab)
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
| FC, WB, E |
|---|---|
| Primary Accession | Q96FC9 |
| Other Accession | NP_689651.1, NP_004390.3 |
| Reactivity | Human |
| Host | Rabbit |
| Clonality | Polyclonal |
| Isotype | Rabbit IgG |
| Calculated MW | 108313 Da |
| Antigen Region | 819-847 aa |
| Gene ID | 1663 |
|---|---|
| Other Names | Probable ATP-dependent RNA helicase DDX11, CHL1-related protein 1, hCHLR1, DEAD/H box protein 11, Keratinocyte growth factor-regulated gene 2 protein, KRG-2, DDX11, CHL1, CHLR1, KRG2 |
| Target/Specificity | This DDX11 antibody is generated from rabbits immunized with a KLH conjugated synthetic peptide between 819-847 amino acids from the C-terminal region of human DDX11. |
| Dilution | WB~~1:1000 FC~~1:10~50 |
| Format | Purified polyclonal antibody supplied in PBS with 0.09% (W/V) sodium azide. This antibody is purified through a protein A column, followed by peptide affinity purification. |
| Storage | Maintain refrigerated at 2-8°C for up to 2 weeks. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | DDX11 Antibody (C-term) is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | DDX11 (HGNC:2736) |
|---|---|
| Function | DNA-dependent ATPase and ATP-dependent DNA helicase that participates in various functions in genomic stability, including DNA replication, DNA repair and heterochromatin organization as well as in ribosomal RNA synthesis (PubMed:10648783, PubMed:21854770, PubMed:23797032, PubMed:26089203, PubMed:26503245). Its double-stranded DNA helicase activity requires either a minimal 5'-single-stranded tail length of approximately 15 nt (flap substrates) or 10 nt length single- stranded gapped DNA substrates of a partial duplex DNA structure for helicase loading and translocation along DNA in a 5' to 3' direction (PubMed:18499658, PubMed:22102414). The helicase activity is capable of displacing duplex regions up to 100 bp, which can be extended up to 500 bp by the replication protein A (RPA) or the cohesion CTF18-replication factor C (Ctf18-RFC) complex activities (PubMed:18499658). Shows also ATPase- and helicase activities on substrates that mimic key DNA intermediates of replication, repair and homologous recombination reactions, including forked duplex, anti-parallel G-quadruplex and three-stranded D-loop DNA molecules (PubMed:22102414, PubMed:26503245). Plays a role in DNA double-strand break (DSB) repair at the DNA replication fork during DNA replication recovery from DNA damage (PubMed:23797032). Recruited with TIMELESS factor upon DNA-replication stress response at DNA replication fork to preserve replication fork progression, and hence ensure DNA replication fidelity (PubMed:26503245). Cooperates also with TIMELESS factor during DNA replication to regulate proper sister chromatid cohesion and mitotic chromosome segregation (PubMed:17105772, PubMed:18499658, PubMed:20124417, PubMed:23116066, PubMed:23797032). Stimulates 5'- single-stranded DNA flap endonuclease activity of FEN1 in an ATP- and helicase-independent manner; and hence it may contribute in Okazaki fragment processing at DNA replication fork during lagging strand DNA synthesis (PubMed:18499658). Its ability to function at DNA replication fork is modulated by its binding to long non-coding RNA (lncRNA) cohesion regulator non-coding RNA DDX11-AS1/CONCR, which is able to increase both DDX11 ATPase activity and binding to DNA replicating regions (PubMed:27477908). Also plays a role in heterochromatin organization (PubMed:21854770). Involved in rRNA transcription activation through binding to active hypomethylated rDNA gene loci by recruiting UBTF and the RNA polymerase Pol I transcriptional machinery (PubMed:26089203). Plays a role in embryonic development and prevention of aneuploidy (By similarity). Involved in melanoma cell proliferation and survival (PubMed:23116066). Associates with chromatin at DNA replication fork regions (PubMed:27477908). Binds to single- and double-stranded DNAs (PubMed:9013641, PubMed:18499658, PubMed:22102414). |
| Cellular Location | Nucleus. Nucleus, nucleolus. Cytoplasm, cytoskeleton, spindle pole. Midbody Cytoplasm, cytoskeleton, microtubule organizing center, centrosome. Note=During the early stages of mitosis, localizes to condensed chromatin and is released from the chromatin with progression to metaphase. Also localizes to the spindle poles throughout mitosis and at the midbody at later stages of mitosis (metaphase to telophase) (PubMed:17105772). In interphase, colocalizes with nucleolin in the nucleolus (PubMed:26089203) |
| Tissue Location | Expressed in melanoma cells. Not detected in epidermal melanocytes of normal skin (at protein level) (PubMed:23116066). Highly expressed in spleen, B-cells, thymus, testis, ovary, small intestine and pancreas (PubMed:9013641). Very low expression seen in brain (PubMed:9013641). Expressed in dividing cells and/or cells undergoing high levels of recombination (PubMed:9013641) No expression detected in cells signaled to terminally differentiate (PubMed:9013641). Expressed weakly in keratinocytes (PubMed:8798685) |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
Background
DEAD box proteins, characterized by the conserved motif Asp-Glu-Ala-Asp (DEAD), are putative RNA helicases. They are implicated in a number of cellular processes involving alteration of RNA secondary structure such as translation initiation, nuclear and mitochondrial splicing, and ribosome and spliceosome assembly. Based on their distribution patterns, some members of this family are believed to be involved in embryogenesis, spermatogenesis, and cellular growth and division. This gene encodes a DEAD box protein, which is an enzyme that possesses both ATPase and DNA helicase activities. This gene is a homolog of the yeast CHL1 gene, and may function to maintain chromosome transmission fidelity and genome stability. Alternative splicing results in multiple transcript variants encoding distinct isoforms.
References
Leman, A.R., et al. J. Cell. Sci. 123 (PT 5), 660-670 (2010) :
Farina, A., et al. J. Biol. Chem. 283(30):20925-20936(2008)
Parish, J.L., et al. Mol. Cell 24(6):867-876(2006)
Parish, J.L., et al. J. Cell. Sci. 119 (PT 23), 4857-4865 (2006) :
Vasa-Nicotera, M., et al. Am. J. Hum. Genet. 76(1):147-151(2005)
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.













 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.