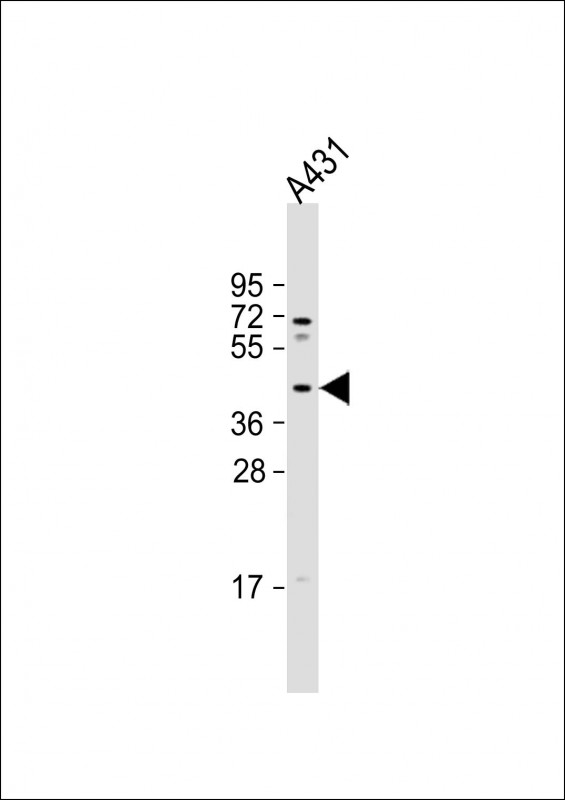
MUL1 Antibody (C-term)
Affinity Purified Rabbit Polyclonal Antibody (Pab)
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
| WB, E |
|---|---|
| Primary Accession | Q969V5 |
| Other Accession | Q4R7G8, NP_078820.2 |
| Reactivity | Human |
| Predicted | Monkey |
| Host | Rabbit |
| Clonality | Polyclonal |
| Isotype | Rabbit IgG |
| Calculated MW | 39800 Da |
| Antigen Region | 272-301 aa |
| Gene ID | 79594 |
|---|---|
| Other Names | Mitochondrial ubiquitin ligase activator of NFKB 1, 632-, E3 SUMO-protein ligase MUL1, E3 ubiquitin-protein ligase MUL1, Growth inhibition and death E3 ligase, Mitochondrial-anchored protein ligase, MAPL, Putative NF-kappa-B-activating protein 266, RING finger protein 218, MUL1, C1orf166, GIDE, MAPL, MULAN, RNF218 |
| Target/Specificity | This MUL1 antibody is generated from rabbits immunized with a KLH conjugated synthetic peptide between 272-301 amino acids from the C-terminal region of human MUL1. |
| Dilution | WB~~1:1000 |
| Format | Purified polyclonal antibody supplied in PBS with 0.09% (W/V) sodium azide. This antibody is purified through a protein A column, followed by peptide affinity purification. |
| Storage | Maintain refrigerated at 2-8°C for up to 2 weeks. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | MUL1 Antibody (C-term) is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | MUL1 |
|---|---|
| Synonyms | C1orf166, GIDE, MAPL, MULAN, RNF218 |
| Function | Exhibits weak E3 ubiquitin-protein ligase activity (PubMed:18591963, PubMed:19407830, PubMed:22410793). E3 ubiquitin ligases accept ubiquitin from an E2 ubiquitin-conjugating enzyme in the form of a thioester and then directly transfer the ubiquitin to targeted substrates (PubMed:18591963, PubMed:19407830, PubMed:22410793). Can ubiquitinate AKT1 preferentially at 'Lys-284' involving 'Lys-48'-linked polyubiquitination and seems to be involved in regulation of Akt signaling by targeting phosphorylated Akt to proteasomal degradation (PubMed:22410793). Mediates polyubiquitination of cytoplasmic TP53 at 'Lys-24' which targets TP53 for proteasomal degradation, thus reducing TP53 levels in the cytoplasm and mitochondrion (PubMed:21597459). Proposed to preferentially act as a SUMO E3 ligase at physiological concentrations (PubMed:19407830). Plays a role in the control of mitochondrial morphology by promoting mitochondrial fragmentation, and influences mitochondrial localization (PubMed:19407830, PubMed:18207745, PubMed:18213395). Likely to promote mitochondrial fission through negatively regulating the mitochondrial fusion proteins MFN1 and MFN2, acting in a pathway that is parallel to the PRKN/PINK1 regulatory pathway (PubMed:24898855). May also be involved in the sumoylation of the membrane fission protein DNM1L (PubMed:18207745, PubMed:19407830). Inhibits cell growth (PubMed:18591963, PubMed:22410793). When overexpressed, activates JNK through MAP3K7/TAK1 and induces caspase-dependent apoptosis (PubMed:23399697). Involved in the modulation of innate immune defense against viruses by inhibiting RIGI-dependent antiviral response (PubMed:23399697). Can mediate RIGI sumoylation and disrupt its polyubiquitination (PubMed:23399697). |
| Cellular Location | Mitochondrion outer membrane; Multi-pass membrane protein. Peroxisome. Note=Transported in mitochondrion- derived vesicles from the mitochondrion to the peroxisome |
| Tissue Location | Widely expressed with highest levels in the heart, skeletal muscle, placenta, kidney and liver. Barely detectable in colon and thymus. |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
Background
E3 ubiquitin-protein ligase that plays a role in the control of mitochondrial morphology. Promotes mitochondrial fragmentation and influences mitochondrial localization. Inhibits cell growth. When overexpressed, activates JNK through MAP3K7/TAK1 and induces caspase-dependent apoptosis. E3 ubiquitin ligases accept ubiquitin from an E2 ubiquitin-conjugating enzyme in the form of a thioester and then directly transfer the ubiquitin to targeted substrates.
References
Laure, L., et al. FEBS J. 277(20):4322-4337(2010)
Braschi, E., et al. EMBO Rep. 10(7):748-754(2009)
Venkatesan, K., et al. Nat. Methods 6(1):83-90(2009)
Zhang, B., et al. Cell Res. 18(9):900-910(2008)
Zhang, H., et al. Biochem. Biophys. Res. Commun. 366(4):898-904(2008)
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.













 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.