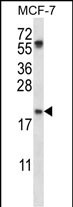
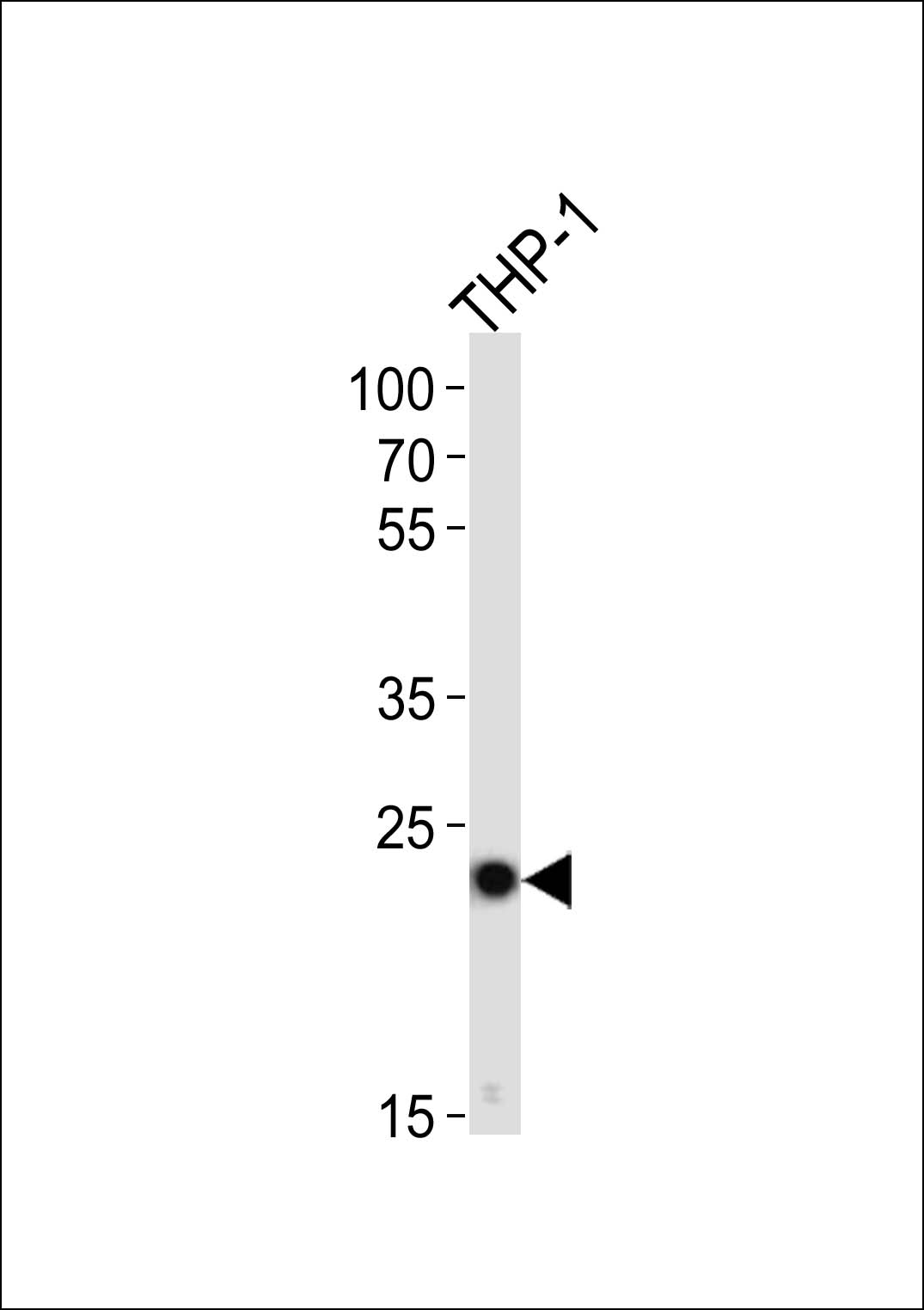
PYCARD Antibody (C-term)
Affinity Purified Rabbit Polyclonal Antibody (Pab)
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
| WB, E |
|---|---|
| Primary Accession | Q9ULZ3 |
| Other Accession | NP_037390.2, NP_660183.1 |
| Reactivity | Human |
| Host | Rabbit |
| Clonality | Polyclonal |
| Isotype | Rabbit IgG |
| Calculated MW | 21627 Da |
| Antigen Region | 129-158 aa |
| Gene ID | 29108 |
|---|---|
| Other Names | Apoptosis-associated speck-like protein containing a CARD, hASC, Caspase recruitment domain-containing protein 5, PYD and CARD domain-containing protein, Target of methylation-induced silencing 1, PYCARD, ASC, CARD5, TMS1 |
| Target/Specificity | This PYCARD antibody is generated from rabbits immunized with a KLH conjugated synthetic peptide between 129-158 amino acids from the C-terminal region of human PYCARD. |
| Dilution | WB~~1:1000 |
| Format | Purified polyclonal antibody supplied in PBS with 0.09% (W/V) sodium azide. This antibody is purified through a protein A column, followed by peptide affinity purification. |
| Storage | Maintain refrigerated at 2-8°C for up to 2 weeks. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | PYCARD Antibody (C-term) is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | PYCARD {ECO:0000303|Ref.4, ECO:0000312|HGNC:HGNC:16608} |
|---|---|
| Function | Functions as a key mediator in apoptosis and inflammation (PubMed:17599095, PubMed:25847972, PubMed:19494289, PubMed:15030775, PubMed:17349957, PubMed:19158675, PubMed:19158676, PubMed:30674671, PubMed:34678144, PubMed:24630722, PubMed:21487011, PubMed:19234215, PubMed:11103777, PubMed:12646168). Promotes caspase-mediated apoptosis involving predominantly caspase-8 and also caspase-9 in a probable cell type-specific manner (PubMed:11103777, PubMed:12646168). Involved in activation of the mitochondrial apoptotic pathway, promotes caspase-8- dependent proteolytic maturation of BID independently of FADD in certain cell types and also mediates mitochondrial translocation of BAX and activates BAX-dependent apoptosis coupled to activation of caspase- 9, -2 and -3 (PubMed:16964285, PubMed:14730312). Involved in innate immune response by acting as an integral adapter in the assembly of various inflammasomes (NLRP1, NLRP2, NLRP3, NLRP6, AIM2 and probably IFI16) which recruit and activate caspase-1 leading to processing and secretion of pro-inflammatory cytokines (PubMed:17599095, PubMed:25847972, PubMed:15030775, PubMed:17349957, PubMed:19158675, PubMed:19158676, PubMed:30674671, PubMed:34678144, PubMed:16982856, PubMed:24630722, PubMed:21487011, PubMed:19234215, PubMed:23530044, PubMed:29440442, PubMed:33980849). Caspase-1-dependent inflammation leads to macrophage pyroptosis, a form of cell death (PubMed:24630722). The function as activating adapter in different types of inflammasomes is mediated by the pyrin and CARD domains and their homotypic interactions (PubMed:19234215, PubMed:14499617, PubMed:24630722). Clustered PYCARD nucleates the formation of caspase-1 filaments through the interaction of their respective CARD domains, acting as a platform for of caspase-1 polymerization (PubMed:24630722). In the NLRP1 and NLRC4 inflammasomes seems not be required but facilitates the processing of procaspase-1 (PubMed:17349957). In cooperation with NOD2 involved in an inflammasome activated by bacterial muramyl dipeptide leading to caspase-1 activation (PubMed:16964285). May be involved in RIGI-triggered pro-inflammatory responses and inflammasome activation (PubMed:19915568). In collaboration with AIM2 which detects cytosolic double-stranded DNA may also be involved in a caspase-1-independent cell death that involves caspase-8 (PubMed:19158675, PubMed:19158676). In adaptive immunity may be involved in maturation of dendritic cells to stimulate T-cell immunity and in cytoskeletal rearrangements coupled to chemotaxis and antigen uptake may be involved in post- transcriptional regulation of the guanine nucleotide exchange factor DOCK2; the latter function is proposed to involve the nuclear form (PubMed:22732093). Also involved in transcriptional activation of cytokines and chemokines independent of the inflammasome; this function may involve AP-1, NF-kappa-B, MAPK and caspase-8 signaling pathways (PubMed:12486103, PubMed:16585594). For regulation of NF-kappa-B activating and inhibiting functions have been reported (PubMed:12486103). Modulates NF-kappa-B induction at the level of the IKK complex by inhibiting kinase activity of CHUK and IKBK (PubMed:12486103, PubMed:16585594). Proposed to compete with RIPK2 for association with CASP1 thereby down-regulating CASP1-mediated RIPK2- dependent NF-kappa-B activation and activating interleukin-1 beta processing (PubMed:16585594). Modulates host resistance to DNA virus infection, probably by inducing the cleavage of and inactivating CGAS in presence of cytoplasmic double-stranded DNA (PubMed:28314590). |
| Cellular Location | Cytoplasm. Inflammasome. Endoplasmic reticulum. Mitochondrion. Nucleus Note=Upstream of caspase activation, a redistribution from the cytoplasm to the aggregates occurs. These appear as hollow, perinuclear spherical, ball-like structures (PubMed:11103777, PubMed:12191486, PubMed:15030775). Upon NLRP3 inflammasome activation redistributes to the perinuclear space localizing to endoplasmic reticulum and mitochondria (PubMed:12191486, PubMed:15030775). Localized primarily to the nucleus in resting monocytes/macrophages and rapidly redistributed to the cytoplasm upon pathogen infection (PubMed:19234215). Localized to large cytoplasmic aggregate appearing as a speck containing AIM2, PYCARD, CASP8 and bacterial DNA after infection with Francisella tularensis (By similarity). {ECO:0000250|UniProtKB:Q9EPB4, ECO:0000269|PubMed:11103777, ECO:0000269|PubMed:12191486, ECO:0000269|PubMed:15030775, ECO:0000269|PubMed:19234215} |
| Tissue Location | Widely expressed at low levels. Detected in peripheral blood leukocytes, lung, small intestine, spleen, thymus, colon and at lower levels in placenta, liver and kidney. Very low expression in skeletal muscle, heart and brain. Expressed in lung epithelial cells (at protein level) (PubMed:23229815). Detected in the leukemia cell lines HL-60 and U-937, but not in Jurkat T-cell lymphoma and Daudi Burkitt's lymphoma. Detected in the melanoma cell line WM35, but not in WM793. Not detected in HeLa cervical carcinoma cells and MOLT-4 lymphocytic leukemia cells. |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
Background
This gene encodes an adaptor protein that is composed of two protein-protein interaction domains: a N-terminal PYRIN-PAAD-DAPIN domain (PYD) and a C-terminal caspase-recruitment domain (CARD). The PYD and CARD domains are members of the six-helix bundle death domain-fold superfamily that mediates assembly of large signaling complexes in the inflammatory and apoptotic signaling pathways via the activation of caspase. In normal cells, this protein is localized to the cytoplasm; however, in cells undergoing apoptosis, it forms ball-like aggregates near the nuclear periphery. Two transcript variants encoding different isoforms have been found for this gene.
References
Grau, E., et al. J. Cancer Res. Clin. Oncol. 136(9):1415-1421(2010)
Motani, K., et al. Cancer Sci. 101(8):1822-1827(2010)
Mishra, B.B., et al. Cell. Microbiol. 12(8):1046-1063(2010)
Cheng, J., et al. J. Cell. Physiol. 222(3):738-747(2010)
McElvania Tekippe, E., et al. PLoS ONE 5 (8), E12320 (2010) :
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.













 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.