PLEKHM1 Antibody
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
| WB, IHC-P, IF, E |
|---|---|
| Primary Accession | Q9Y4G2 |
| Other Accession | Q9Y4G2, 160419247 |
| Reactivity | Human |
| Host | Rabbit |
| Clonality | Polyclonal |
| Isotype | IgG |
| Calculated MW | 117443 Da |
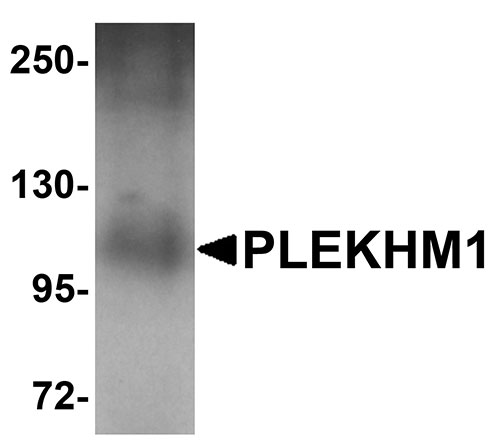
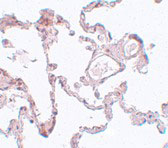
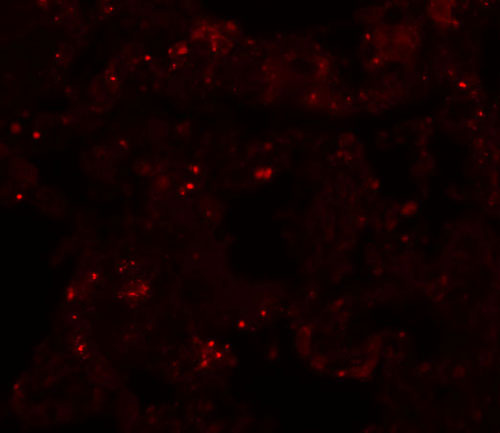
| Application Notes | PLEKHM1 antibody can be used for detection of PLEKHM1 by Western blot at 1 µg/mL. Antibody can also be used for immunohistochemistry starting at 5 µg/mL. For immunofluorescence start at 20 µg/mL. |
| Gene ID | 9842 |
|---|---|
| Target/Specificity | PLEKHM1; |
| Reconstitution & Storage | PLEKHM1 antibody can be stored at 4℃ for three months and -20℃, stable for up to one year. As with all antibodies care should be taken to avoid repeated freeze thaw cycles. Antibodies should not be exposed to prolonged high temperatures. |
| Precautions | PLEKHM1 Antibody is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | PLEKHM1 (HGNC:29017) |
|---|---|
| Synonyms | KIAA0356 |
| Function | Acts as a multivalent adapter protein that regulates Rab7- dependent and HOPS complex-dependent fusion events in the endolysosomal system and couples autophagic and the endocytic trafficking pathways. Acts as a dual effector of RAB7A and ARL8B that simultaneously binds these GTPases, bringing about clustering and fusion of late endosomes and lysosomes (PubMed:25498145, PubMed:28325809). Required for late stages of endolysosomal maturation, facilitating both endocytosis- mediated degradation of growth factor receptors and autophagosome clearance. Interaction with Arl8b is a crucial factor in the terminal maturation of autophagosomes and to mediate autophagosome-lysosome fusion (PubMed:25498145). Positively regulates lysosome peripheral distribution and ruffled border formation in osteoclasts (By similarity). May be involved in negative regulation of endocytic transport from early endosome to late endosome/lysosome implicating its association with Rab7 (PubMed:20943950). May have a role in sialyl-lex- mediated transduction of apoptotic signals (PubMed:12820725). Involved in bone resorption (By similarity). |
| Cellular Location | Autolysosome membrane. Endosome membrane. Late endosome membrane. Lysosome membrane. Note=In case of infection colocalizes with Salmonella typhimurium sifA in proximity of Salmonella-containing vacuole (SCV) (PubMed:25500191). |
| Tissue Location | Expressed in placenta, liver, prostate, thymus, spleen, ovary, colon, colon carcinoma and peripheral blood lymphocytes (PBL). Weakly expressed in brain, lung, kidney, and testis. No expression in heart, skeletal muscle, pancreas and small intestine Predominantly expressed in the breast carcinoma cell line MCF-7 |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
Background
PLEKHM1 Antibody: PLEKHM1 is a member of the M family of Pleckstrin homolog domain-containing proteins, a group of proteins containing a RUN domain, two pleckstrin homology domains, and a cysteine-rich domain. It was identified through segregation analysis as a cause of osteopetrosis in humans. PLEKHM1 co-localizes with Rab7 to late endosomal/lysosomal vesicles in HEK293 and osteoclast-like cells, with this co-localization dependent on the prenylation of Rab7. Monocytes from a patient homozygous for a mutated form of PLEKHM1differentiated into osteoclasts normally, but failed to form ruffled borders and showed little evidence of bone resorbtion when cultured on dentine discs. Another mutation of PLEKHM1 impaired vesicular acidification and increased TRACP secretion in osteoclasts, suggesting that PLEKHM1 has critical roles in endosomal maturation and may be important in osteoclast-osteoblast cross-talk.
References
Van Wesenbeeck L, Odgren PR, Mackay CA, et al. Localization of the gene causing the osteopetrotic phenotype in the incisors absent (ia) rat on chromosome 10q32.1. J. Bone Miner. Res.2004; 19:183-9.
Van Wesenbeeck L, Odgren PR, Coxon FP, et al. Involvement of PLEKHM1 in osteoclastic vesicular transport and osteopetrosis in incisors absent rats and humans. J. Clin. Invest.2007; 117:919-30.
Del Fattore A, Fornari R, Van Wesenbeeck L, et al. A new heterozygous mutation (R714C) of the osteopetrosis gene, pleckstrin homolog domain containing family M (with run domain) member 1 (PLEKHM1), impairs vesicular acidification and increases TRACP secretion in osteoclasts. J. Bone Miner. Res.2008; 23:380-91.
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.













 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.