ZBTB3 Antibody
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
| WB, IHC-P, IF, E |
|---|---|
| Primary Accession | Q9H5J0 |
| Other Accession | AAH25249, 13376146 |
| Reactivity | Human, Mouse, Rat |
| Host | Rabbit |
| Clonality | Polyclonal |
| Isotype | IgG |
| Calculated MW | 61827 Da |
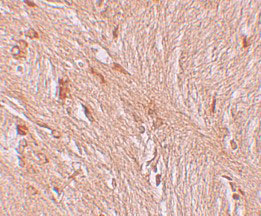
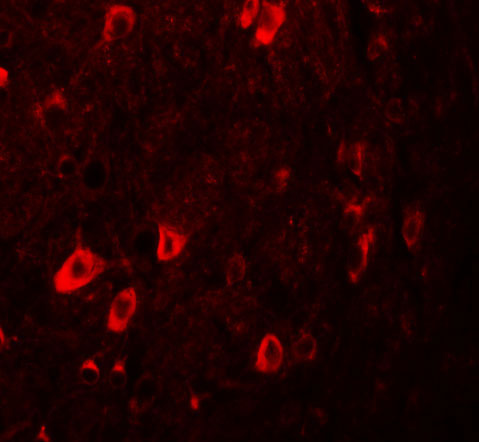
| Application Notes | ZBTB3 antibody can be used for detection of ZBTB3 by Western blot at 1 - 2 µg/mL. Antibody can also be used for immunohistochemistry starting at 2.5 µg/mL. For immunofluorescence start at 20 µg/mL. |
| Gene ID | 79842 |
|---|---|
| Target/Specificity | ZBTB3; At least three isoforms of ZBTB3 are known to exist. This antibody is predicted to not cross-react with other ZBTB protein family members. |
| Reconstitution & Storage | ZBTB3 antibody can be stored at 4℃ for three months and -20℃, stable for up to one year. As with all antibodies care should be taken to avoid repeated freeze thaw cycles. Antibodies should not be exposed to prolonged high temperatures. |
| Precautions | ZBTB3 Antibody is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | ZBTB3 |
|---|---|
| Function | May be involved in transcriptional regulation. |
| Cellular Location | Nucleus. |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
Background
ZBTB3 Antibody: The ZBTB family of proteins is comprised of diverse zinc finger proteins that also contain a BTB (BR-C, ttk and bab) domain. While little is known about ZBTB3, the related protein ZBTB2 is thought to be phosphorylated in response to the DNA damage, probably by either ATM or ATR. Other ZBTB proteins, such as ZBTB4 and ZBTB38 bind methylated DNA and repress transcription, suggesting that ZBTB3 may also act as a transcription repressor.
References
Strausberg RL, Feingold EA, Grouse LH, et al. Generation and initial analysis of more than 15,000 full-length human and mouse cDNA sequences. Proc. Natl. Acad. Sci. USA2002; 99:16899-903.
Matsuoka S, Ballif BA, Smogorzewska A, et al. ATM and ATR substrate analysis reveals extensive protein networks responsive to DNA damage. Science2007; 1160-1166.
Filion GJP, Zhenilo S, Salozhin S, et al. A family of zinc finger proteins that bind methylated DNA and repress transcription. Mol. Cell. Biol.2006; 26:169-81.
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.













 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.