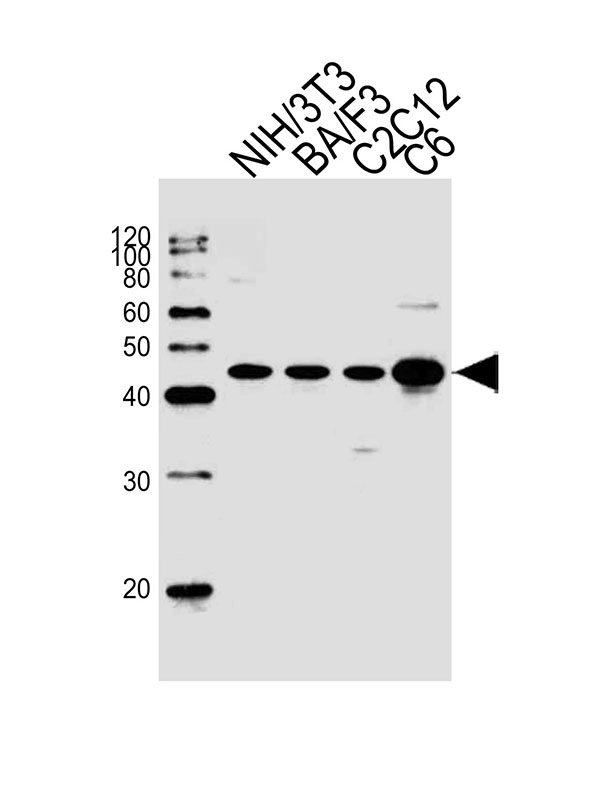
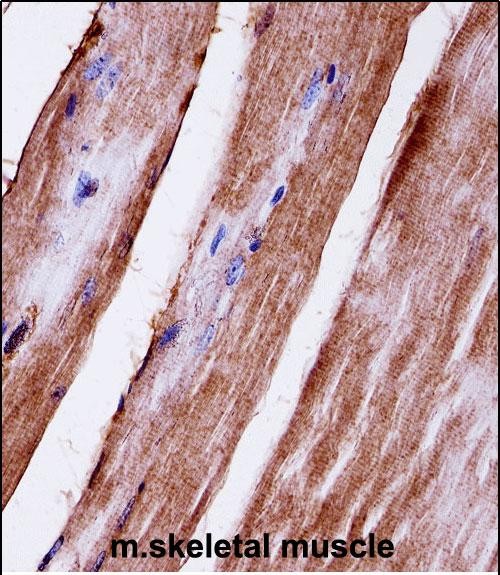
Mouse Mapk3 Antibody (C-term)
Affinity Purified Rabbit Polyclonal Antibody (Pab)
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
| FC, IHC-P, WB |
|---|---|
| Primary Accession | Q63844 |
| Other Accession | NP_036082.1 |
| Reactivity | Mouse, Rat |
| Predicted | Human |
| Host | Rabbit |
| Clonality | Polyclonal |
| Calculated MW | H=43;M=43;Rat=43 KDa |
| Isotype | Rabbit IgG |
| Antigen Source | MOUSE |
| Gene ID | 26417 |
|---|---|
| Antigen Region | 353-380 aa |
| Other Names | Mapk3; Erk1; Prkm3; Mitogen-activated protein kinase 3; ERT2; Extracellular signal-regulated kinase 1; Insulin-stimulated MAP2 kinase; MAP kinase isoform p44; MNK1; Microtubule-associated protein 2 kinase; p44-ERK1 |
| Dilution | WB~~1:1000 IHC-P~~1:10~50 FC~~1:25 |
| Target/Specificity | This Mouse Mapk3 antibody is generated from rabbits immunized with a KLH conjugated synthetic peptide between 353-380 amino acids from the C-terminal region of mouse Mapk3. |
| Format | Purified polyclonal antibody supplied in PBS with 0.09% (W/V) sodium azide. This antibody is purified through a protein A column, followed by peptide affinity purification. |
| Storage | Maintain refrigerated at 2-8°C for up to 2 weeks. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | Mouse Mapk3 Antibody (C-term) is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | Mapk3 |
|---|---|
| Synonyms | Erk1, Prkm3 |
| Function | Serine/threonine kinase which acts as an essential component of the MAP kinase signal transduction pathway. MAPK1/ERK2 and MAPK3/ERK1 are the 2 MAPKs which play an important role in the MAPK/ERK cascade. They participate also in a signaling cascade initiated by activated KIT and KITLG/SCF. Depending on the cellular context, the MAPK/ERK cascade mediates diverse biological functions such as cell growth, adhesion, survival and differentiation through the regulation of transcription, translation, cytoskeletal rearrangements. The MAPK/ERK cascade also plays a role in initiation and regulation of meiosis, mitosis, and postmitotic functions in differentiated cells by phosphorylating a number of transcription factors. About 160 substrates have already been discovered for ERKs. Many of these substrates are localized in the nucleus, and seem to participate in the regulation of transcription upon stimulation. However, other substrates are found in the cytosol as well as in other cellular organelles, and those are responsible for processes such as translation, mitosis and apoptosis. Moreover, the MAPK/ERK cascade is also involved in the regulation of the endosomal dynamics, including lysosome processing and endosome cycling through the perinuclear recycling compartment (PNRC); as well as in the fragmentation of the Golgi apparatus during mitosis. The substrates include transcription factors (such as ATF2, BCL6, ELK1, ERF, FOS, HSF4 or SPZ1), cytoskeletal elements (such as CANX, CTTN, GJA1, MAP2, MAPT, PXN, SORBS3 or STMN1), regulators of apoptosis (such as BAD, BTG2, CASP9, DAPK1, IER3, MCL1 or PPARG), regulators of translation (such as EIF4EBP1) and a variety of other signaling-related molecules (like ARHGEF2, DEPTOR, FRS2 or GRB10). Protein kinases (such as RAF1, RPS6KA1/RSK1, RPS6KA3/RSK2, RPS6KA2/RSK3, RPS6KA6/RSK4, SYK, MKNK1/MNK1, MKNK2/MNK2, RPS6KA5/MSK1, RPS6KA4/MSK2, MAPKAPK3 or MAPKAPK5) and phosphatases (such as DUSP1, DUSP4, DUSP6 or DUSP16) are other substrates which enable the propagation the MAPK/ERK signal to additional cytosolic and nuclear targets, thereby extending the specificity of the cascade. |
| Cellular Location | Cytoplasm {ECO:0000250|UniProtKB:P21708}. Nucleus. Membrane, caveola {ECO:0000250|UniProtKB:P21708}. Cell junction, focal adhesion Note=Autophosphorylation at Thr-207 promotes nuclear localization (By similarity). PEA15-binding redirects the biological outcome of MAPK3 kinase-signaling by sequestering MAPK3 into the cytoplasm (PubMed:11702783). {ECO:0000250|UniProtKB:P27361, ECO:0000269|PubMed:11702783} |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
Background
Involved in both the initiation and regulation of meiosis, mitosis, and postmitotic functions in differentiated cells by phosphorylating a number of transcription factors such as ELK-1. Phosphorylates EIF4EBP1; required for initiation of translation. Phosphorylates microtubule-associated protein 2 (MAP2) and heat shock factor protein 4 (HSF4) (By similarity). Phosphorylates SPZ1.
References
Chen, W., et al. Biochem. Biophys. Res. Commun. 401(3):339-343(2010)
Chandrakesan, P., et al. J. Biol. Chem. 285(43):33485-33498(2010)
Alter, B.J., et al. J. Neurosci. 30(34):11537-11547(2010)
Spruce, T., et al. Dev. Cell 19(2):207-219(2010)
Bremer, J., et al. PLoS ONE 5 (8), E12450 (2010) :
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.













 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.