PIN1 Antibody
Mouse Monoclonal Antibody (Mab)
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
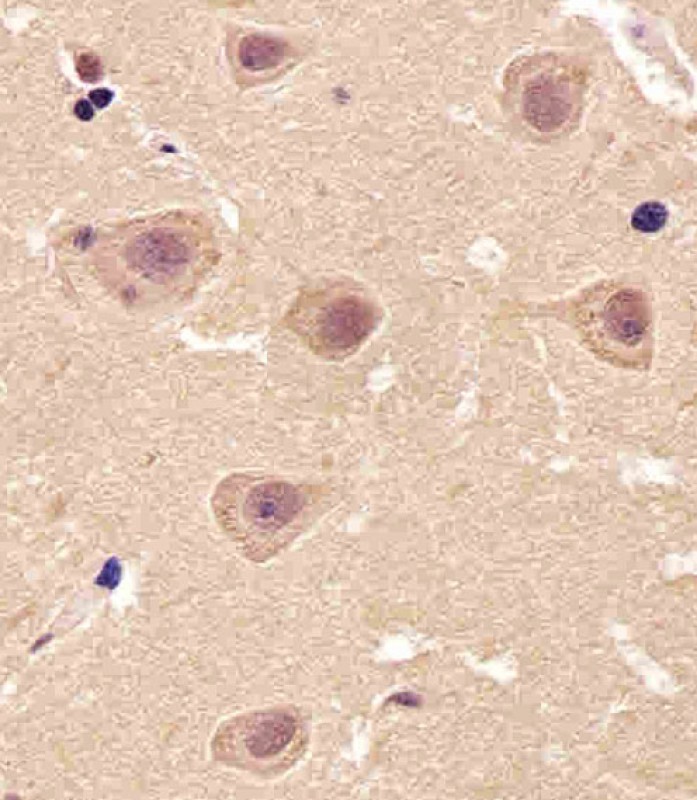
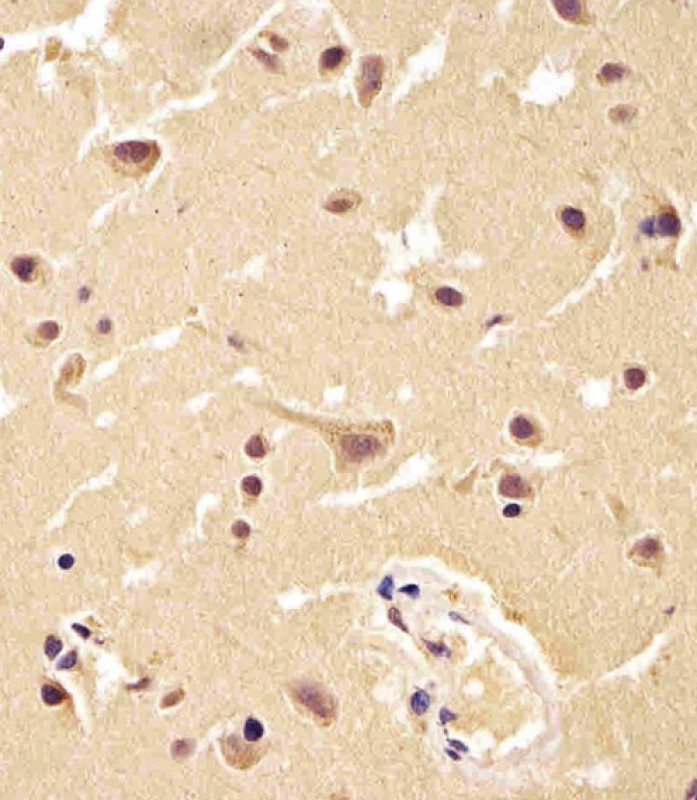
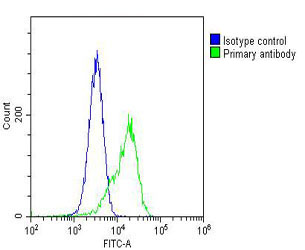
| WB, IHC-P, IHC, FC |
|---|---|
| Primary Accession | Q13526 |
| Reactivity | Human, Mouse |
| Predicted | Rat |
| Host | Mouse |
| Clonality | Monoclonal |
| Calculated MW | H=18;M=18;Rat=18 KDa |
| Isotype | IgG1 |
| Antigen Source | Human |
| Gene ID | 5300 |
|---|---|
| Antigen Region | 1-143 aa |
| Other Names | PIN1;Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1; Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1; Peptidyl-prolyl cis-trans isomerase Pin1; Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1; Rotamase Pin1 |
| Dilution | WB~~1:1000 IHC~~1:25 IHC-P~~1:25 FC~~1:25 |
| Target/Specificity | Purified His-tagged PIN1 protein was used to produced this monoclonal antibody. |
| Format | Purified monoclonal antibody supplied in PBS with 0.09% (W/V) sodium azide. This antibody is purified through a protein G column, followed by dialysis against PBS. |
| Storage | Maintain refrigerated at 2-8°C for up to 2 weeks. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | PIN1 Antibody is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | PIN1 |
|---|---|
| Function | Peptidyl-prolyl cis/trans isomerase (PPIase) that binds to and isomerizes specific phosphorylated Ser/Thr-Pro (pSer/Thr-Pro) motifs (PubMed:21497122, PubMed:23623683, PubMed:29686383). By inducing conformational changes in a subset of phosphorylated proteins, acts as a molecular switch in multiple cellular processes (PubMed:21497122, PubMed:22033920, PubMed:23623683). Displays a preference for acidic residues located N-terminally to the proline bond to be isomerized. Regulates mitosis presumably by interacting with NIMA and attenuating its mitosis-promoting activity. Down-regulates kinase activity of BTK (PubMed:16644721). Can transactivate multiple oncogenes and induce centrosome amplification, chromosome instability and cell transformation. Required for the efficient dephosphorylation and recycling of RAF1 after mitogen activation (PubMed:15664191). Binds and targets PML and BCL6 for degradation in a phosphorylation-dependent manner (PubMed:17828269). Acts as a regulator of JNK cascade by binding to phosphorylated FBXW7, disrupting FBXW7 dimerization and promoting FBXW7 autoubiquitination and degradation: degradation of FBXW7 leads to subsequent stabilization of JUN (PubMed:22608923). May facilitate the ubiquitination and proteasomal degradation of RBBP8/CtIP through CUL3/KLHL15 E3 ubiquitin-protein ligase complex, hence favors DNA double-strand repair through error-prone non-homologous end joining (NHEJ) over error-free, RBBP8-mediated homologous recombination (HR) (PubMed:23623683, PubMed:27561354). Upon IL33-induced lung inflammation, catalyzes cis-trans isomerization of phosphorylated IRAK3/IRAK-M, inducing IRAK3 stabilization, nuclear translocation and expression of pro-inflammatory genes in dendritic cells (PubMed:29686383). |
| Cellular Location | Nucleus. Nucleus speckle. Cytoplasm Note=Colocalizes with NEK6 in the nucleus (PubMed:16476580). Mainly localized in the nucleus but phosphorylation at Ser-71 by DAPK1 results in inhibition of its nuclear localization (PubMed:21497122) |
| Tissue Location | Expressed in immune cells in the lung (at protein level) (PubMed:29686383). The phosphorylated form at Ser-71 is expressed in normal breast tissue cells but not in breast cancer cells |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
Background
Essential PPIase that regulates mitosis presumably by interacting with NIMA and attenuating its mitosis-promoting activity. Displays a preference for an acidic residue N-terminal to the isomerized proline bond. Catalyzes pSer/Thr-Pro cis/trans isomerizations. Down-regulates kinase activity of BTK. Can transactivate multiple oncogenes and induce centrosome amplification, chromosome instability and cell transformation. Required for the efficient dephosphorylation and recycling of RAF1 after mitogen activation.
References
Ebert L., et al. Submitted (MAY-2004) to the EMBL/GenBank/DDBJ databases.
Lu K.P., et al. Nature 380:544-547(1996).
Kalnine N., et al. Submitted (OCT-2004) to the EMBL/GenBank/DDBJ databases.
Ota T., et al. Nat. Genet. 36:40-45(2004).
Mural R.J., et al. Submitted (JUL-2005) to the EMBL/GenBank/DDBJ databases.
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.













 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.