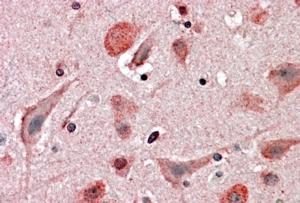
Goat Anti-HIP2 Antibody
Peptide-affinity purified goat antibody
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
| WB, IHC, E |
|---|---|
| Primary Accession | P61086 |
| Other Accession | NP_001104583, 3093 |
| Reactivity | Human |
| Predicted | Mouse, Pig, Dog, Cow |
| Host | Goat |
| Clonality | Polyclonal |
| Concentration | 100ug/200ul |
| Isotype | IgG |
| Calculated MW | 22407 Da |
| Gene ID | 3093 |
|---|---|
| Other Names | Ubiquitin-conjugating enzyme E2 K, 6.3.2.19, Huntingtin-interacting protein 2, HIP-2, Ubiquitin carrier protein, Ubiquitin-conjugating enzyme E2-25 kDa, Ubiquitin-conjugating enzyme E2(25K), Ubiquitin-conjugating enzyme E2-25K, Ubiquitin-protein ligase, UBE2K, HIP2, LIG |
| Format | 0.5 mg IgG/ml in Tris saline (20mM Tris pH7.3, 150mM NaCl), 0.02% sodium azide, with 0.5% bovine serum albumin |
| Storage | Maintain refrigerated at 2-8°C for up to 6 months. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | Goat Anti-HIP2 Antibody is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | UBE2K |
|---|---|
| Synonyms | HIP2, LIG |
| Function | Accepts ubiquitin from the E1 complex and catalyzes its covalent attachment to other proteins. In vitro, in the presence or in the absence of BRCA1-BARD1 E3 ubiquitin-protein ligase complex, catalyzes the synthesis of 'Lys-48'-linked polyubiquitin chains. Does not transfer ubiquitin directly to but elongates monoubiquitinated substrate protein. Mediates the selective degradation of short-lived and abnormal proteins, such as the endoplasmic reticulum-associated degradation (ERAD) of misfolded lumenal proteins. Ubiquitinates huntingtin. May mediate foam cell formation by the suppression of apoptosis of lipid-bearing macrophages through ubiquitination and subsequence degradation of p53/TP53. Proposed to be involved in ubiquitination and proteolytic processing of NF-kappa-B; in vitro supports ubiquitination of NFKB1. In case of infection by cytomegaloviruses may be involved in the US11-dependent degradation of MHC class I heavy chains following their export from the ER to the cytosol. In case of viral infections may be involved in the HPV E7 protein-dependent degradation of RB1. |
| Cellular Location | Cytoplasm {ECO:0000250|UniProtKB:P61085}. |
| Tissue Location | Expressed in all tissues tested, including spleen, thymus, prostate, testis, ovary, small intestine, colon, peripheral blood leukocytes, T-lymphocytes, monocytes, granulocytes and bone marrow mononuclear cells. Highly expressed in brain, with highest levels found in cortex and striatum and at lower levels in cerebellum and brainstem. |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
Background
The protein encoded by this gene belongs to the ubiquitin-conjugating enzyme family. This protein interacts with RING finger proteins, and it can ubiquitinate huntingtin, the gene product for Huntington's disease. Known functions for this protein include a role in aggregate formation of expanded polyglutamine proteins and the suppression of apoptosis in polyglutamine diseases, a role in the dislocation of newly synthesized MHC class I heavy chains from the endoplasmic reticulum, and involvement in foam cell formation. Multiple transcript variants encoding different isoforms have been identified for this gene.
References
Hip2 interacts with and destabilizes Smac/DIABLO. Bae Y, et al. Biochem Biophys Res Commun, 2010 Jul 9. PMID 20537984.
Ubc9 sumoylation regulates SUMO target discrimination. Knipscheer P, et al. Mol Cell, 2008 Aug 8. PMID 18691969.
E2-BRCA1 RING interactions dictate synthesis of mono- or specific polyubiquitin chain linkages. Christensen DE, et al. Nat Struct Mol Biol, 2007 Oct. PMID 17873885.
UbcH8 regulates ubiquitin and ISG15 conjugation to RIG-I. Arimoto K, et al. Mol Immunol, 2008 Feb. PMID 17719635.
Large-scale mapping of human protein-protein interactions by mass spectrometry. Ewing RM, et al. Mol Syst Biol, 2007. PMID 17353931.
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.













 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.