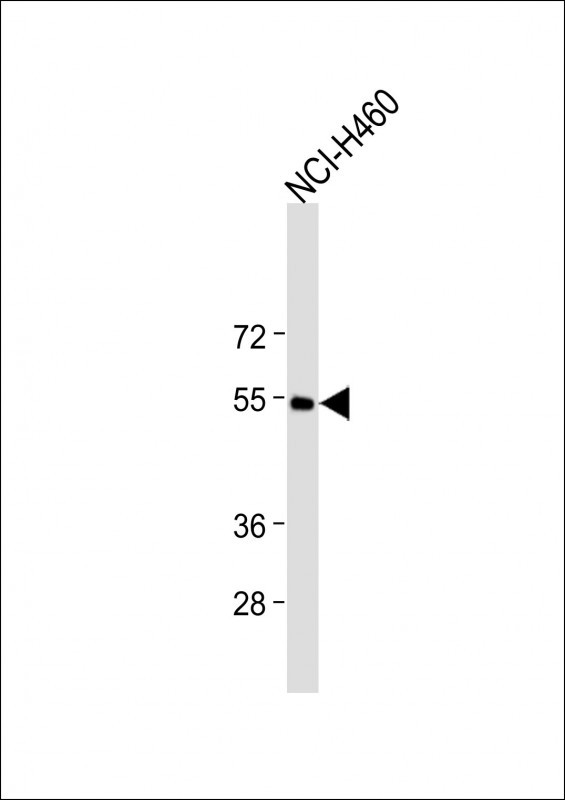
CXCR4 Antibody(N-term)
Affinity Purified Rabbit Polyclonal Antibody (Pab)
- SPECIFICATION
- CITATIONS: 2
- PROTOCOLS
- BACKGROUND

Application
| WB, E |
|---|---|
| Primary Accession | P61073 |
| Other Accession | O08565, Q764M9, P70658, Q28474, P25930, NP_001008540.1, Q28553 |
| Reactivity | Human |
| Predicted | Bovine, Monkey, Mouse, Pig, Rat, Sheep |
| Host | Rabbit |
| Clonality | Polyclonal |
| Isotype | Rabbit IgG |
| Calculated MW | 39746 Da |
| Antigen Region | 50-79 aa |
| Gene ID | 7852 |
|---|---|
| Other Names | C-X-C chemokine receptor type 4, CXC-R4, CXCR-4, FB22, Fusin, HM89, LCR1, Leukocyte-derived seven transmembrane domain receptor, LESTR, Lipopolysaccharide-associated protein 3, LAP-3, LPS-associated protein 3, NPYRL, Stromal cell-derived factor 1 receptor, SDF-1 receptor, CD184, CXCR4 |
| Target/Specificity | This CXCR4 antibody is generated from rabbits immunized with a KLH conjugated synthetic peptide between 50-79 amino acids from the N-terminal region of human CXCR4. |
| Dilution | WB~~1:1000 |
| Format | Purified polyclonal antibody supplied in PBS with 0.09% (W/V) sodium azide. This antibody is purified through a protein A column, followed by peptide affinity purification. |
| Storage | Maintain refrigerated at 2-8°C for up to 2 weeks. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | CXCR4 Antibody(N-term) is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | CXCR4 |
|---|---|
| Function | Receptor for the C-X-C chemokine CXCL12/SDF-1 that transduces a signal by increasing intracellular calcium ion levels and enhancing MAPK1/MAPK3 activation (PubMed:10452968, PubMed:28978524, PubMed:18799424, PubMed:24912431). Involved in the AKT signaling cascade (PubMed:24912431). Plays a role in regulation of cell migration, e.g. during wound healing (PubMed:28978524). Acts as a receptor for extracellular ubiquitin; leading to enhanced intracellular calcium ions and reduced cellular cAMP levels (PubMed:20228059). Binds bacterial lipopolysaccharide (LPS) et mediates LPS-induced inflammatory response, including TNF secretion by monocytes (PubMed:11276205). Involved in hematopoiesis and in cardiac ventricular septum formation. Also plays an essential role in vascularization of the gastrointestinal tract, probably by regulating vascular branching and/or remodeling processes in endothelial cells. Involved in cerebellar development. In the CNS, could mediate hippocampal-neuron survival (By similarity). |
| Cellular Location | Cell membrane; Multi-pass membrane protein. Cell junction. Early endosome. Late endosome. Lysosome. Note=In unstimulated cells, diffuse pattern on plasma membrane. On agonist stimulation, colocalizes with ITCH at the plasma membrane where it becomes ubiquitinated. In the presence of antigen, distributes to the immunological synapse forming at the T- cell-APC contact area, where it localizes at the peripheral and distal supramolecular activation cluster (SMAC) |
| Tissue Location | Expressed in numerous tissues, such as peripheral blood leukocytes, spleen, thymus, spinal cord, heart, placenta, lung, liver, skeletal muscle, kidney, pancreas, cerebellum, cerebral cortex and medulla (in microglia as well as in astrocytes), brain microvascular, coronary artery and umbilical cord endothelial cells Isoform 1 is predominant in all tissues tested |
- Primary Antibodies
- Cancer
- Immunology
- Metabolism
- Microbiology
- Neuroscience
- Signal Transduction
- Antibody Collections
- GPCR Antibodies
- Druggable G protein-coupled receptors (GPCRs)
- Host cell receptor for virus entry
- Host-virus interaction
- Immune system process
- Chemokine receptor family
- Chemokine signaling
- Inflammation mediated by chemokine and cytokine signaling
- Lysosome
- Intestinal immune network for IgA production

Provided below are standard protocols that you may find useful for product applications.
Background
This gene encodes a CXC chemokine receptor specific for stromal cell-derived factor-1. The protein has 7 transmembrane regions and is located on the cell surface. It acts with the CD4 protein to support HIV entry into cells and is also highly expressed in breast cancer cells. Mutations in this gene have been associated with WHIM (warts, hypogammaglobulinemia, infections, and myelokathexis) syndrome. Alternate transcriptional splice variants, encoding different isoforms, have been characterized. [provided by RefSeq].
References
Hammad, M.M., et al. J. Biol. Chem. 285(45):34653-34664(2010)
Wang, A., et al. Arthritis Rheum. 62(11):3436-3446(2010)
Cheng, M., et al. Circ. Res. 107(9):1083-1093(2010)
Sanematsu, F., et al. Circ. Res. 107(9):1102-1105(2010)
O'Hayre, M., et al. PLoS ONE 5 (7), E11716 (2010) :
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.















 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.