APEX1 Antibody (N-term)
Purified Rabbit Polyclonal Antibody (Pab)
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
| IF, WB, E |
|---|---|
| Primary Accession | P27695 |
| Reactivity | Human |
| Host | Rabbit |
| Clonality | Polyclonal |
| Isotype | Rabbit IgG |
| Calculated MW | 35555 Da |
| Antigen Region | 30-64 aa |
| Gene ID | 328 |
|---|---|
| Other Names | DNA-(apurinic or apyrimidinic site) lyase, 31--, APEX nuclease, APEN, Apurinic-apyrimidinic endonuclease 1, AP endonuclease 1, APE-1, REF-1, Redox factor-1, DNA-(apurinic or apyrimidinic site) lyase, mitochondrial, APEX1, APE, APE1, APEX, APX, HAP1, REF1 |
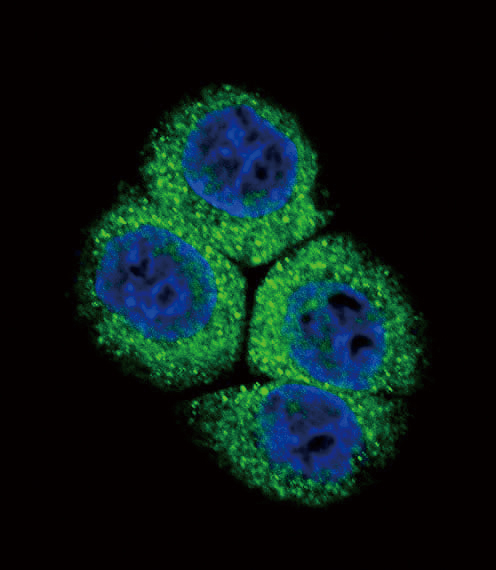
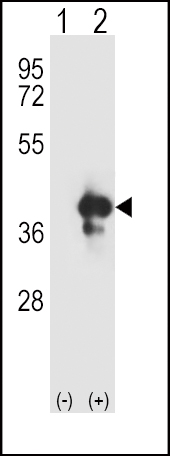
| Target/Specificity | This APEX1 antibody is generated from rabbits immunized with a KLH conjugated synthetic peptide between 30-64 amino acids from the N-terminal region of human APEX1. |
| Dilution | IF~~1:10~50 WB~~1:1000 |
| Format | Purified polyclonal antibody supplied in PBS with 0.09% (W/V) sodium azide. This antibody is prepared by Saturated Ammonium Sulfate (SAS) precipitation followed by dialysis against PBS. |
| Storage | Maintain refrigerated at 2-8°C for up to 2 weeks. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | APEX1 Antibody (N-term) is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | APEX1 |
|---|---|
| Synonyms | APE, APE1, APEX, APX, HAP1, REF1 |
| Function | Multifunctional protein that plays a central role in the cellular response to oxidative stress. The two major activities of APEX1 are DNA repair and redox regulation of transcriptional factors. Functions as an apurinic/apyrimidinic (AP) endodeoxyribonuclease in the DNA base excision repair (BER) pathway of DNA lesions induced by oxidative and alkylating agents. Initiates repair of AP sites in DNA by catalyzing hydrolytic incision of the phosphodiester backbone immediately adjacent to the damage, generating a single-strand break with 5'-deoxyribose phosphate and 3'-hydroxyl ends. Also incises at AP sites in the DNA strand of DNA/RNA hybrids, single-stranded DNA regions of R-loop structures, and single-stranded RNA molecules. Has 3'-5' exoribonuclease activity on mismatched deoxyribonucleotides at the 3' termini of nicked or gapped DNA molecules during short-patch BER. Possesses DNA 3' phosphodiesterase activity capable of removing lesions (such as phosphoglycolate) blocking the 3' side of DNA strand breaks. May also play a role in the epigenetic regulation of gene expression by participating in DNA demethylation. Acts as a loading factor for POLB onto non-incised AP sites in DNA and stimulates the 5'-terminal deoxyribose 5'-phosphate (dRp) excision activity of POLB. Plays a role in protection from granzyme-mediated cellular repair leading to cell death. Also involved in the DNA cleavage step of class switch recombination (CSR). On the other hand, APEX1 also exerts reversible nuclear redox activity to regulate DNA binding affinity and transcriptional activity of transcriptional factors by controlling the redox status of their DNA-binding domain, such as the FOS/JUN AP-1 complex after exposure to IR. Involved in calcium-dependent down- regulation of parathyroid hormone (PTH) expression by binding to negative calcium response elements (nCaREs). Together with HNRNPL or the dimer XRCC5/XRCC6, associates with nCaRE, acting as an activator of transcriptional repression. Stimulates the YBX1-mediated MDR1 promoter activity, when acetylated at Lys-6 and Lys-7, leading to drug resistance. Acts also as an endoribonuclease involved in the control of single-stranded RNA metabolism. Plays a role in regulating MYC mRNA turnover by preferentially cleaving in between UA and CA dinucleotides of the MYC coding region determinant (CRD). In association with NMD1, plays a role in the rRNA quality control process during cell cycle progression. Associates, together with YBX1, on the MDR1 promoter. Together with NPM1, associates with rRNA. Binds DNA and RNA. |
| Cellular Location | Nucleus {ECO:0000255|PROSITE-ProRule:PRU00764}. Nucleus, nucleolus. Nucleus speckle. Endoplasmic reticulum. Cytoplasm Note=Detected in the cytoplasm of B-cells stimulated to switch (By similarity). Colocalized with SIRT1 in the nucleus. Colocalized with YBX1 in nuclear speckles after genotoxic stress. Together with OGG1 is recruited to nuclear speckles in UVA-irradiated cells. Colocalized with nucleolin and NPM1 in the nucleolus. Its nucleolar localization is cell cycle dependent and requires active rRNA transcription. Colocalized with calreticulin in the endoplasmic reticulum. Translocation from the nucleus to the cytoplasm is stimulated in presence of nitric oxide (NO) and function in a CRM1-dependent manner, possibly as a consequence of demasking a nuclear export signal (amino acid position 64-80). S- nitrosylation at Cys-93 and Cys-310 regulates its nuclear-cytosolic shuttling. Ubiquitinated form is localized predominantly in the cytoplasm. |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
Background
Apurinic/apyrimidinic (AP) sites occur frequently in DNA molecules by spontaneous hydrolysis, by DNA damaging agents or by DNA glycosylases that remove specific abnormal bases. AP sites are pre-mutagenic lesions that can prevent normal DNA replication so the cell contains systems to identify and repair such sites. Class II AP endonucleases cleave the phosphodiester backbone 5' to the AP site. This protein is the major AP endonuclease in human cells.
References
Vascotto,C., Mol. Cell. Biol. 29 (7), 1834-1854 (2009)
Li,W.Q., Carcinogenesis 30 (3), 500-505 (2009)
Lo,Y.L., Cancer Epidemiol. Biomarkers Prev. 18 (1), 223-229 (2009)
Bhakat,K.K., EMBO J. 22 (23), 6299-6309 (2003)
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.














 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.