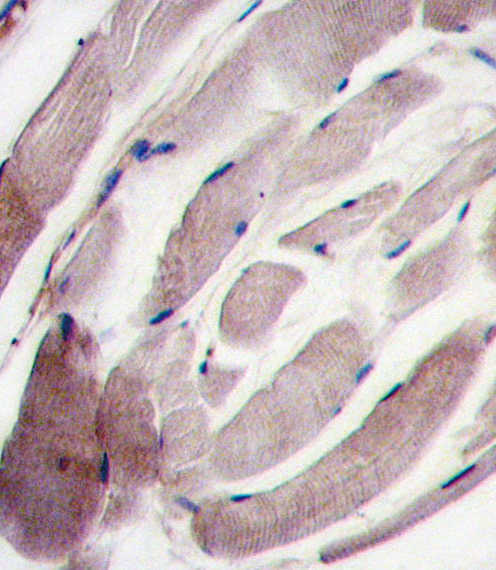
MAPK12 Antibody (C-term)
Purified Rabbit Polyclonal Antibody (Pab)
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
| IHC-P, WB, E |
|---|---|
| Primary Accession | P53778 |
| Reactivity | Human, Mouse |
| Host | Rabbit |
| Clonality | Polyclonal |
| Isotype | Rabbit IgG |
| Calculated MW | 41940 Da |
| Antigen Region | 336-367 aa |
| Gene ID | 6300 |
|---|---|
| Other Names | Mitogen-activated protein kinase 12, MAP kinase 12, MAPK 12, Extracellular signal-regulated kinase 6, ERK-6, Mitogen-activated protein kinase p38 gamma, MAP kinase p38 gamma, Stress-activated protein kinase 3, MAPK12, ERK6, SAPK3 |
| Target/Specificity | This MAPK12 antibody is generated from rabbits immunized with a KLH conjugated synthetic peptide between 336-367 amino acids from the C-terminal region of human MAPK12. |
| Dilution | WB~~1:1000 IHC-P~~1:10~50 |
| Format | Purified polyclonal antibody supplied in PBS with 0.09% (W/V) sodium azide. This antibody is prepared by Saturated Ammonium Sulfate (SAS) precipitation followed by dialysis against PBS. |
| Storage | Maintain refrigerated at 2-8°C for up to 2 weeks. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | MAPK12 Antibody (C-term) is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | MAPK12 |
|---|---|
| Synonyms | ERK6, SAPK3 |
| Function | Serine/threonine kinase which acts as an essential component of the MAP kinase signal transduction pathway. MAPK12 is one of the four p38 MAPKs which play an important role in the cascades of cellular responses evoked by extracellular stimuli such as pro-inflammatory cytokines or physical stress leading to direct activation of transcription factors such as ELK1 and ATF2. Accordingly, p38 MAPKs phosphorylate a broad range of proteins and it has been estimated that they may have approximately 200 to 300 substrates each. Some of the targets are downstream kinases such as MAPKAPK2, which are activated through phosphorylation and further phosphorylate additional targets. Plays a role in myoblast differentiation and also in the down- regulation of cyclin D1 in response to hypoxia in adrenal cells suggesting MAPK12 may inhibit cell proliferation while promoting differentiation. Phosphorylates DLG1. Following osmotic shock, MAPK12 in the cell nucleus increases its association with nuclear DLG1, thereby causing dissociation of DLG1-SFPQ complexes. This function is independent of its catalytic activity and could affect mRNA processing and/or gene transcription to aid cell adaptation to osmolarity changes in the environment. Regulates UV-induced checkpoint signaling and repair of UV-induced DNA damage and G2 arrest after gamma-radiation exposure. MAPK12 is involved in the regulation of SLC2A1 expression and basal glucose uptake in L6 myotubes; and negatively regulates SLC2A4 expression and contraction-mediated glucose uptake in adult skeletal muscle. C-Jun (JUN) phosphorylation is stimulated by MAPK14 and inhibited by MAPK12, leading to a distinct AP-1 regulation. MAPK12 is required for the normal kinetochore localization of PLK1, prevents chromosomal instability and supports mitotic cell viability. MAPK12- signaling is also positively regulating the expansion of transient amplifying myogenic precursor cells during muscle growth and regeneration. |
| Cellular Location | Cytoplasm. Nucleus. Mitochondrion. Note=Mitochondrial when associated with SH3BP5. In skeletal muscle colocalizes with SNTA1 at the neuromuscular junction and throughout the sarcolemma (By similarity). |
| Tissue Location | Highly expressed in skeletal muscle and heart. |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
Background
P38 gamma is a member of the MAP kinase family. MAP kinases act as an integration point for multiple biochemical signals, and are involved in a wide variety of cellular processes such as proliferation, differentiation, transcription regulation and development. Activation of members of the mitogen-activated protein kinase family is a major mechanism for transduction of extracellular signals. Stress-activated protein kinases are one subclass of MAP kinases. P38 gamma functions as a signal transducer during differentiation of myoblasts to myotubes.
References
Garcia-Lora, A., et al., Cancer Immunol. Immunother. 52(1):59-64 (2003).
Julien, C., et al., J. Biol. Chem. 278(43):42615-42624 (2003).
Robinson, M.J., et al., J. Biol. Chem. 277(7):5094-5100 (2002).
Court, N.W., et al., J. Mol. Cell. Cardiol. 34(4):413-426 (2002).
Wang, X., et al., Mol. Cell. Biol. 20(13):4543-4552 (2000).
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.














 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.